Custom TARGATT™ HEK293 Knock-in Cell Line Service Details
ASC provides custom knockin cell line generation service using the TARGATT™ HEK293 Master Cell Line, which is ideal for single gene knockin and uniform, stable expression of your protein*.
- High efficiency & stringent gene knockin: 40% without enrichment and ~90% with enrichment
- Site-specific integration: into the well-defined H11 active, intergenic (safe harbor) locus
- Isogenic knockin cell lines: one variant - one locus - one cell line
- Uniform and high-level gene expression
- Low off-target integration
- Stable gene knockin with unidirectional integration
- Large transgene knockin (up to 20 kb)
- Overcomes challenges associates with random insertion: gene silencing, multiple copy gene integration, ablated gene expression
Standard Deliverables:
- Three clones that are positive for the targeted insertion; with two vials of each clone at 1 x 10^6 cells/vial.
- Milestone update/report; final report with a detailed description of each procedure, including targeting vector design, construction and validation, transfection condition, genotyping strategy, and results.
Standard Workflow and Timeline:
| Service |
|
1. Generation of donor plasmid by cloning the gene of interest into TARGATT™ cloning plasmid
|
| 2. Transfection of the donor and TARGATT™ integrase plasmids into the master cell line |
|
3. Sorting/ enrichment of transfected cells and single cell cloning
|
|
4. Clone confirmation by genotyping
|
|
5. Cell expansion and cryopreservation
|
Timeline: 3-4 months
Applications:
- Generation of Isogenic knockin cell lines for precise comparison of different genes
- Reporter gene insertion
- Gene overexpression
- Protein production
- Directed-evolution of proteins (vaccine development, drug screening, cell-based gene therapy)
- Genome-wide screening
*The TARGATT™ HEK293 Master Cell Line is also suitable for building large mammalian cell libraries, with non-viral high-throughput library screens and fewer cell counts. It combines the scalability, affordability, and ease-of-use of bacterial/ yeast systems and the advantages of using mammalian cells (closer to human environment and post-translational modifications) for efficient and stable gene knockin into cell lines and for library generation.
If your application involves cell library preparation, please contact Applied StemCell for details.
Supporting Material:
TARGATT™ HEK293 Master Cell Line Knock-in System
- The TARGATT™ HEK293 Master Cell Line was generated for fast and site-specific gene knockin in HEK293 cells, using an easy-to-use gene knockin approach. The master cell line contains an “attP” integrase-recognition landing pad engineered into the well-defined hH11 safe harbor locus in the genome which has been shown to express high levels of protein.
- Donor Plasmid: The gene of interest will be cloned into a TARGATT™ cloning plasmid that contains the “attB” sequence, to generate the donor plasmid.
- TARGATT™ integrase plasmid: to catalyze the irreversible integration of the gene of interest into the preselecting docking site in the H11 locus in the TARGATT™ HEK293 Master Cell Line.
The TARGATT™ integrase and the donor plasmid will be transfected into the TARGATT™ HEK293 Master cell line using well-optimized protocols to enable stable integration of the gene of interest into the cell line.
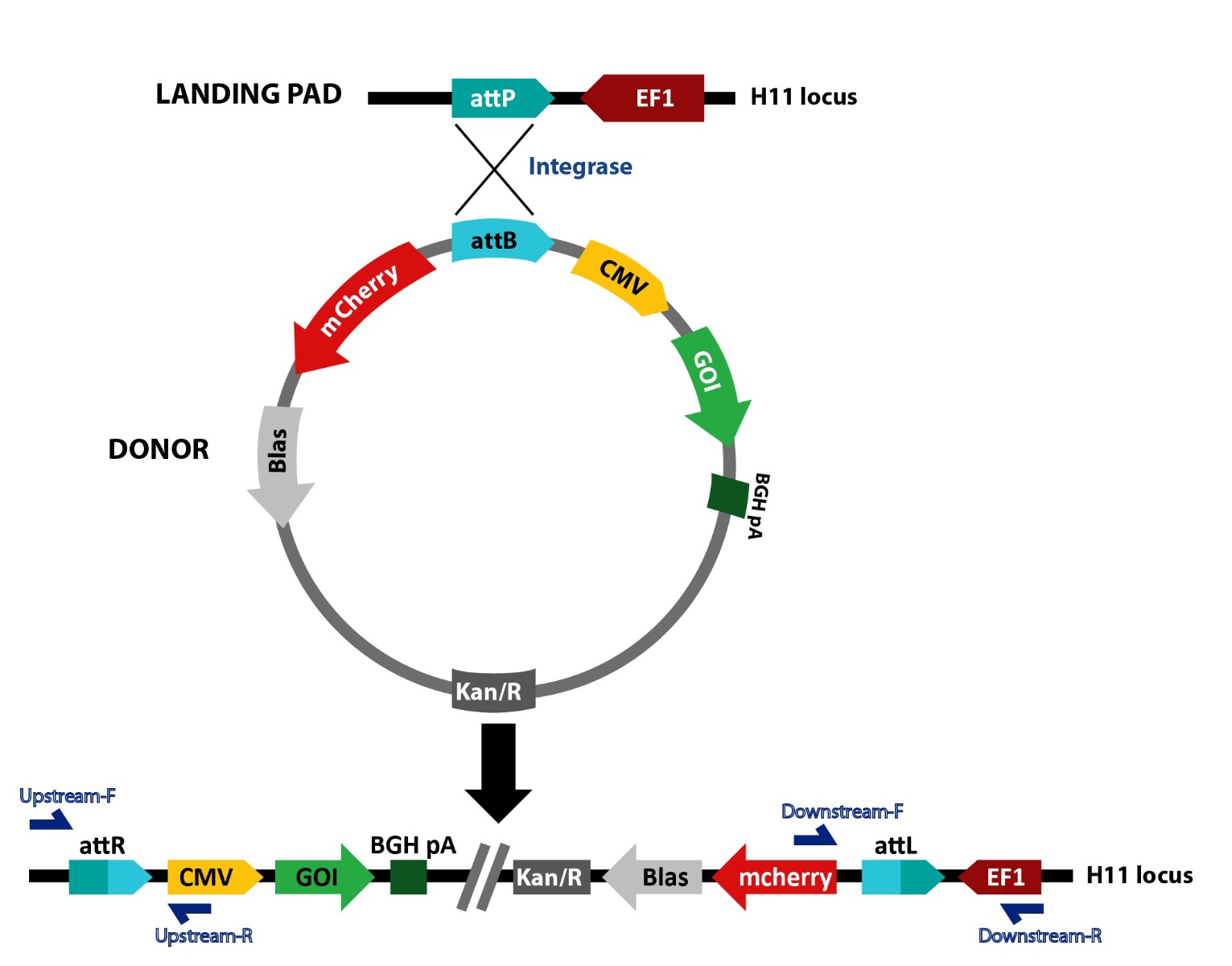
Figure 1. Schematic representation of TARGATT™ site-specific transgene integration mediated by integrase. The TARGATT™ HEK Master Cell Line was engineered with the attP landing pad at the hH11 safe harbor locus. The TARGATT™ plasmid containing the integrase recognition site, attB is used to clone the transgene. The integrase catalyzes an irreversible reaction between the attP site in the genome and attB site in the donor vector, resulting in integration of the gene of interest at the selected H11 locus. The cells containing the gene of interest can be enriched using the selection marker (gray box).
The TARGATT™ HEK293 Master cell line and integrase system enables site-specific and very efficient DNA integration (>40% without enrichment and ~90% with enrichment) without disruption of internal genes.
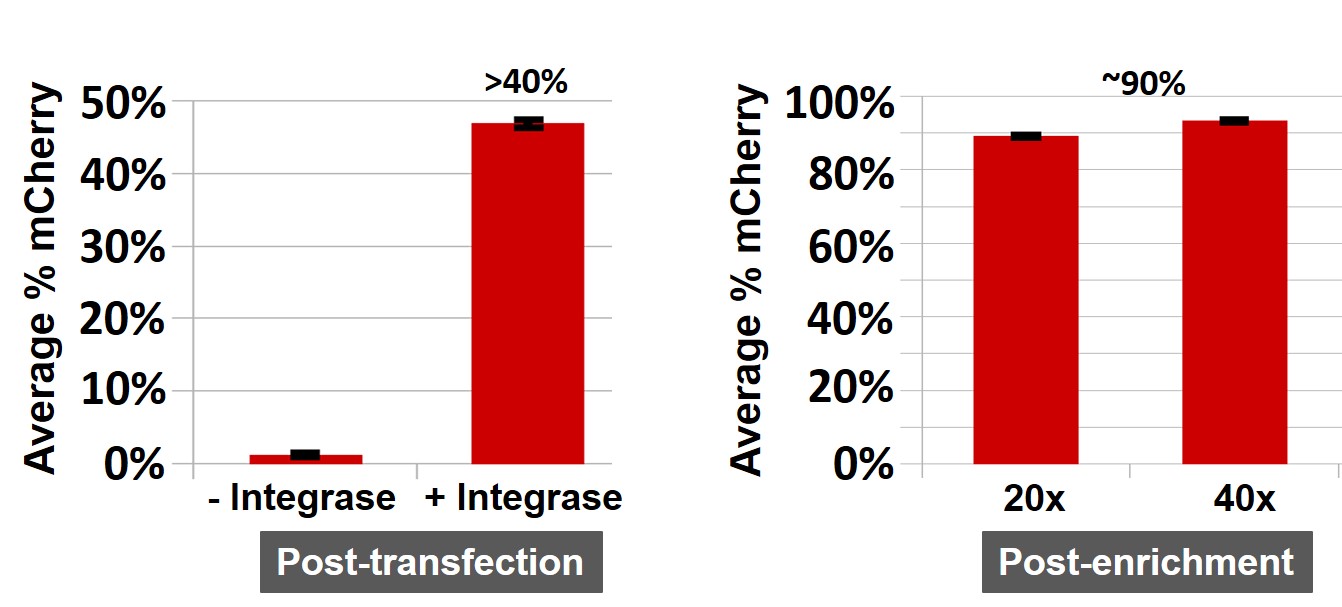
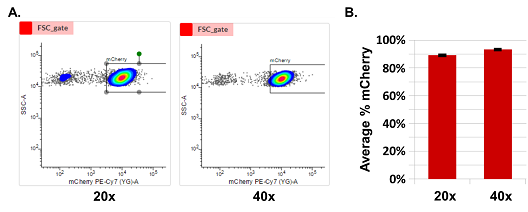
Figure 3. The mCherry integration into the TARGATT™ HEK293 master cell line. Left: Integration mediated by the integrase 72 hours post-transfection. Cells were transfected with the mCherry positive control plasmid and either the provided TARGATT™ integrase plasmid (+Integrase) or a mutant TARGATT™ integrase plasmid deficient for integration (-Integrase). The mCherry plasmid has no promoter and requires the ubiquitous EF1 promoter in the landing pad after integration to express the reporter gene. The integration efficiency of mCherry knockin into landing pad was >40%, without selection. Right:Blasticidin enrichment of TARGATT™ HEK293 cells with a knocked-in mCherry-blasticidin plasmid. Cell pools (with 20x and 40x split ratio) were enriched in selection medium for 3 weeks (without cell sorting). The enrichment of mCherry was about 90% after blasticidin selection. Data represents the mean SE of two representative experiments done in triplicates
TARGATT™ Knockin Cell Lines
A Knockin cell line is one in which a foreign gene or DNA fragment (transgene) is inserted into a cell line genome. These knockin cell lines make powerful research tools for various applications including reporter gene insertion, gene overexpression, and bioproduction. Traditionally, knockin cell lines were generated by random insertion of the foreign genes which carries with it several disadvantages including repeat-induced gene silencing, disruption of internal genes, and genomic instability. More recently, the CRISPR-Cas9 technology has become very popular and is widely used to engineer knockout and knockin cell lines. While CRISPR genome editing is very adaptable and versatile for the generation cell lines with a variety of genetic modifications in a site-specific manner and at a desired locus, it is not very efficient for the integration of large transgenes.
TARGATT™ Technology for site-specific gene knockin:
Applied StemCell’s (ASC) proprietary, TARGATT™ technology enables site-specific integration of large DNA fragments more efficiently. The TARGATT™ technology is based on serine integrases which catalyze efficient, site-specific DNA cleavage, DNA strand exchange and ligation between an integrase-recognition “attP” sequence in the genome (docking site or landing pad) and a corresponding “attB” sequence on the donor vector, resulting in stable gene integration. The landing pad can be engineered into a pre-selected, intergenic, transcriptionally active genomic locus (safe harbor locus) to ensure stable, uniform gene expression.
ASC’s TARGATT™ integrase technology is a very versatile gene knockin tool and it has been successfully adapted in cell lines, mouse, rat, pigs and rabbits models.
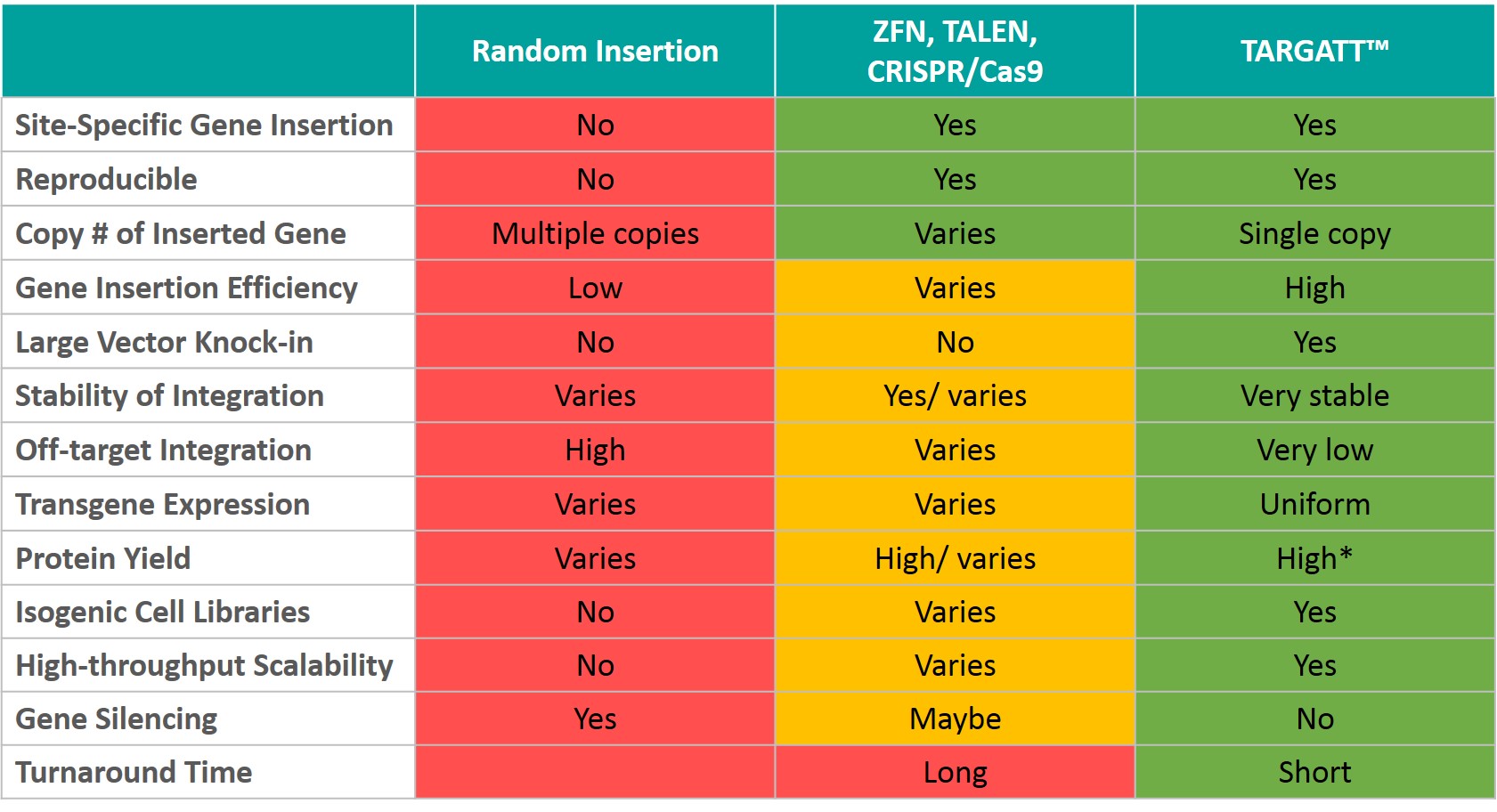
Comparing TARGATT™ and existing gene editing technologies for generating stable knockin cell lines:
How does the TARGATT™ technology compare to CRISPR/Cas9 or nuclease-based technologies?
Nuclease-based genome engineering technologies such as TALENs, ZFNs and especially CRISPR/Cas9 have revolutionized biomedical research in the past decade. CRISPR/Cas9 genome editing is very adaptable and versatile enabling the generation of a cell line and animal models with a variety of genetic modifications such as gene knockout, knockin, conditional knockout, and more, in a site-specific manner and at a desired locus (genetic or safe harbor).
However, CRISPR/Cas9 efficiency for transgene knockin is limited by the size of the transgene. In contrast, while the efficiency of knockin using TARGATT™ is very high and not limited by size of the transgene, the TARGATT™ technology requires an integrase-recognition landing pad (docking site) to be engineered into the desired locus (preferably a safe harbor locus).
While the engineering of a landing is an additional step before gene integration, Applied StemCell has successfully generated “master” cell lines, mouse, rat, pig, and rabbit models and have successfully used them for transgenic cell line and animal modeling, for basic and applied research applications. We have several peer-reviewed journal publications citing the use of TARGATT™ technology for various applications.