Newsletter
CASE STUDY: Heterozygous Large Deletion of a Targeted Sequence in Human iPSCs
Goal: To delete a large fragment of DNA (~100kb) from a specific locus in control human iPSCs to mimic a chromosomal abnormality seen in a rare neurodevelopmental disorder.
1. Human iPSCs were obtained from the customer and the cell line was validated for single cell clonability and other parameters.

Figure 1. Schematic representation of the workflow undertaken for the project.
2. Guide RNAs (gRNAs) were designed based on the distance to the modification site and the off-target profile. The gRNAs were cloned into Cas9-expressing plasmids and validated for indel formation in iPSCs. Two gRNAs, upstream and one downstream of targeted regions, were identified after subsequent analysis of the cells by PCR and sequencing to confirm indel formation at the gRNA cutting site.
3. The 2 gRNAs were then co-transfected into iPSCs and single cell clones were isolated. Genomic DNA from the single cell clones was subjected to PCR (figure 2 and 3) using primer pairs for upstream and downstream of the targeted sequence, and verified by Sanger sequencing (not shown). One positive heterozygous clone was identified with the deletion of the ~100 kb sequence at targeted chromosomal locus.
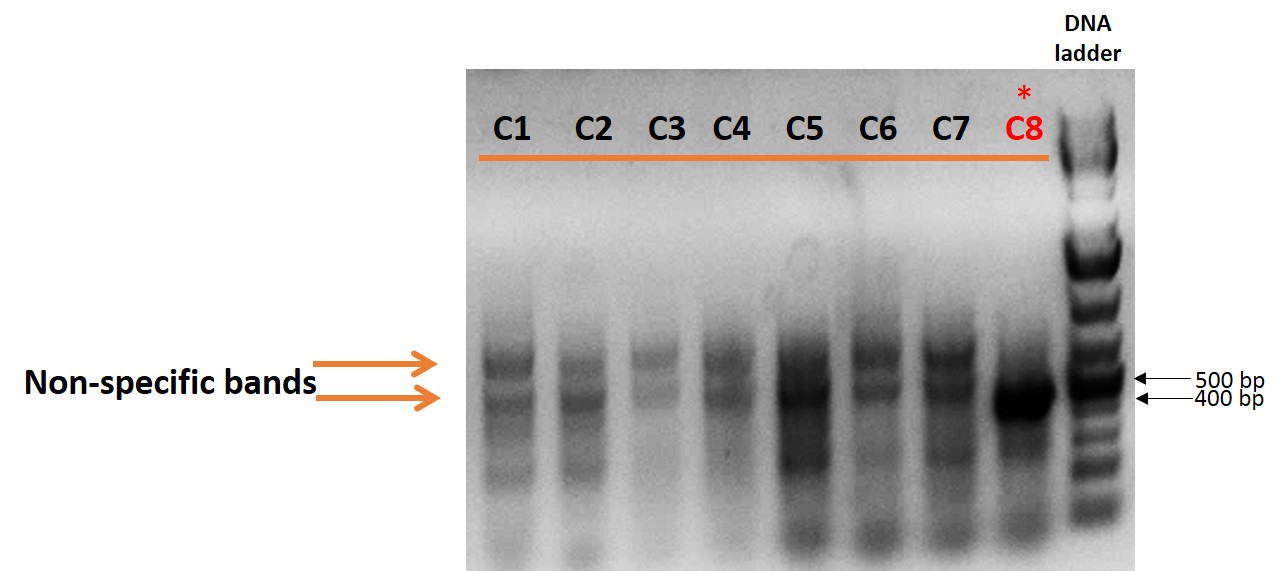
Figure 2. PCR gel electrophoresis to detect knockout of ~100kb fragment in a specific chromosomal locus in human iPSCs, showed a shorter band at the size of 440bp with clone C8. Further sequencing identified this clone (C8) to be positive (*) for the targeted deletion.
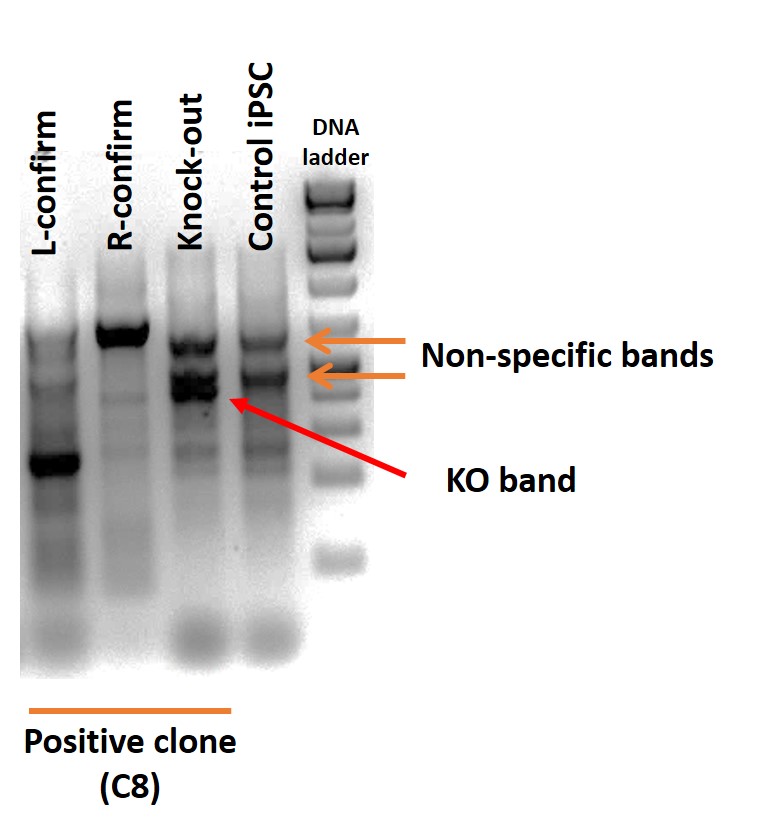
Figure 3. PCR to confirm deletion and to detect homozygous/heterozygous clone with primers flanking the locus of interest: L confirm: upstream forward/reverse primers; R confirm: downstream forward/reverse primers. Clone C8 shows bands for both upstream and downstream PCRs, suggesting that clone C8 is a heterozygous clone.

