Newsletter
CRISPR Knockin Mouse Model Generation
We use an affordable, optimized CRISPR/Cas9 custom knock-in mouse model generation service that allows us to offer a diverse portfolio of knock-in modifications, gene-specific knockin (up to 6 kb), and safe harbor locus knockin. ASC's comprehensive knock-in service allows for the insertion of humanized genes, reporter genes, tags, and more. We work with you to design a specific project outline that captures all of your mouse knock-in model requirements.
- Locus specific (or) safe harbor locus knock-in (e.g., Rosa26, H11, others)
- Knock-in reporter genes/tags, human gene/exons, fusion genes, replacement, and more
- Safe harbor knock-in: CRISPR (Rosa26, H11, AAVS1)
- Efficient CRISPR strategies and protocols for large fragment knock-in
- F1 breeding for germline transmission
- Novel CRISPR strategy for higher efficiency and success rate
- Animal generated in an AAALAC-accredited animal facility
- Animal IP belongs to customers
- Fast turnaround of 5-8 months
- Free consultation
- We also offer a repository of off-shelf mouse models
Products and Services
Case Studies
Case Study 1
CRISPR Knock-in Model - Generation of site-specific 2 kb large fragment knock-in mouse using CRISPR/Cas9
Goal: To insert a 2 kb large fragment DNA (gene of interest) at “a specified locus” in the mouse genome using CRISPR/Cas9.
The project was designed using a well optimized protocol to generate the transgenic mice: (1) Cas9 mRNA and gRNA were produced by in vitro transcription; (2) donor vector was constructed by in-fusion method: the plasmid contained 1 kb long 5’ and 1 kb long 3’ homologous arms flanking the gene of interest (2kb); (3) the mixture of Cas9 mRNA, gRNA and donor vector was microinjected into fertilized eggs of C57BL/6j mouse background.
Using a panel of genotyping primer pairs, three out of 31 pups born after microinjection (#15, 19, and 26) were identified as founders (F0), with the gene of interest inserted at the desired locus.
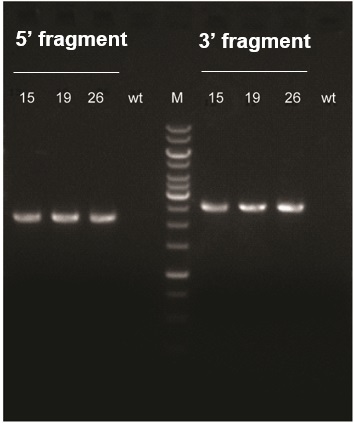
Figure 1: Agarose gel electrophoresis of PCR results in F0 mice (#15, 19, and 26) with site-specific gene knock-in. The left part of the gel shows the 5’ junction fragment (2,191 bp), and the right part of the gel shows the 3’ junction fragment (2,557 bp). [wt: wildtype control; M: 1 kb DNA ladder].
Case Study 2
CRISPR Knock-in Model - Site-specific knock-in of a 27 bp tag in mice using CRISPR/Cas9
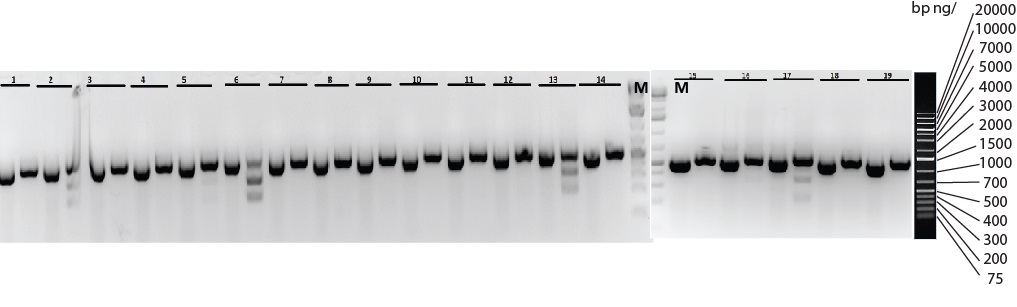
Figure 1: Five (#2, 5, 6 13 and 17) out of nineteen pups, 26%, were genotyped by restriction enzyme digest and sequencing to confirm the correct knock-in on one of the alleles (heterozygotes). A sequence with desired knock-in generated a different cutting pattern compared to its wild type counterpart.
Case Study 3
CRISPR/Cas9 Knock-in Mouse Model
Cdkn2a-e(Luc-2A-tdTomato-2A-CreERT2-WPRE-pA)1 KI mouse model
Knock-in Strategy
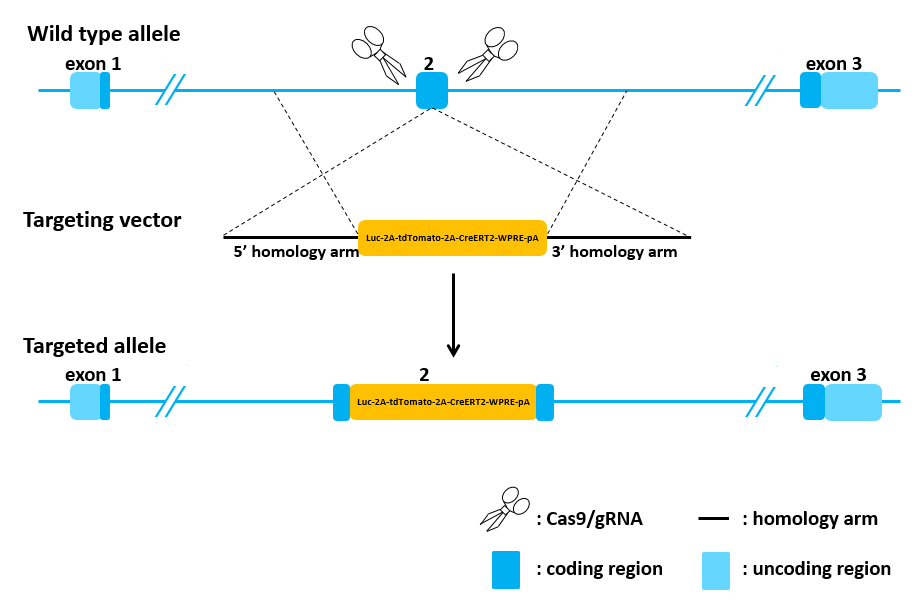
Figure 1. Targeting strategy for Luc-2A-tdTomato-2A-CreERT2-WPRE-pA knock-in
Long PCR genotyping of homologous recombinant F0 mice
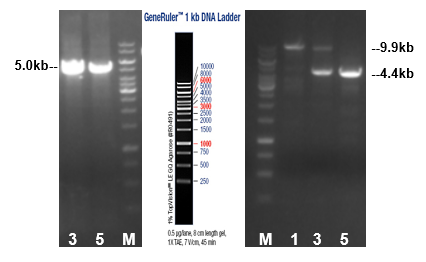
Figure 2. Agarose gel electrophoresis of PCR products. NO.3, 5 were the positive homologous recombinant F0 mice; 5arm: 5’homologous recombinant fragment. 3arm: 3’homologous recombinant fragment. (number: the number of F0 mice; M: 1kb DNA ladder)



