Newsletter
Gene Fusion in iPSCs with CRISPR: Translocation, Inversion & Deletion
Chromosomal translocations and deletions that result in the re-arrangement of separate genes to form chimeric “fusion” genes comprise the most common mutation class in many types of cancer. While fusion genes have been used as markers for cancer diagnosis and prognosis, and for anti-cancer therapies, their functional role in cancer development and progression is largely unknown. The iPS cell lines can provide the necessary research tools to understand the genetic mechanisms involved in cancer development.
Applied StemCell is a unique iPSC service provider that can genetically modify iPSCs with complex, clinically relevant chromosomal rearrangements (translocation, inversion, deletion, and insertion) and further differentiate the iPSCs to the cell lineage-of-choice to better understand the development of cancer and its response to drugs.
ASC Advantages
- High success rate
- We can genetically modify your healthy or diseased iPSCs; control lines are available
- Single cell cloning (clonal isolation)
- Pluripotency maintained throughout the genome editing process
- GMP iPSC Gene Editing Available >> Learn More
GMP iPSC Gene Editing & Additional Services Available >> Learn More
Products and Services
Case Studies
Gene Fusion in Master iPSC Line, ASE-9211
As an expert in engineering/ correcting mutations in control/disease iPSC genome editing, Applied StemCell’s superior CRISPR designs and scientific team also handle complex projects that go well beyond the normal modifications (gene knockout, point mutation/ SNV, reporter/tag knock-in, large transgene knock-in) that customers require.
We can handle complex projects that involve multiplexed genome editing as well as one of our unique service highlights that includes engineering gene fusion mutations (translocation, inversion, deletion, insertion) using CRISPR/Cas9.
Below is a quick snapshot of a project where our scientists successfully engineered a gene translocation mutation in human iPSC line, ASE-9211.
Gene Fusion Research Using iPSCs
Chromosomal translocation is a type of gene fusion where a segment of one chromosome breaks and reattaches to a different location within the same chromosome or to another chromosome. The anaplastic lymphoma kinase (ALK) which is a receptor-type protein tyrosine kinase on chromosome 2, is frequently involved in chromosomal translocations and is indicated in 3-7% of non-small cell lung cancer (NSCLC). Most ALK fusions are caused by the genomic rearrangement of intron 19 of ALK which results in the ALK kinase domain at the 3’ region of the fusion transcript with a partner gene on the 5’ end. One such ALK fusion involves the partner gene KIF5B, a member of the kinesin family of proteins located on chromosome 10. In one variant of the KIF5B-ALK fusions, the genomic breakpoint is located on intron 24 of KIF5B which includes the coiled-coil domain of KIF5B and intron 19 of ALK (COSF1058) which includes the whole ALK tyrosine kinase domain. It is hypothesized that the dimerization promoted by the coiled-coil domain of KIF5B along with its ubiquitous promoter would aberrantly activate the kinase activity of ALK thus contributing to oncogenesis in NSCLC.
Here, our scientists at Applied StemCell engineered the above mentioned KIF5B[24]-ALK[19] gene fusion in a human iPSC line, ASE-9211 using CRISPR/Cas9 and proprietary design algorithms.
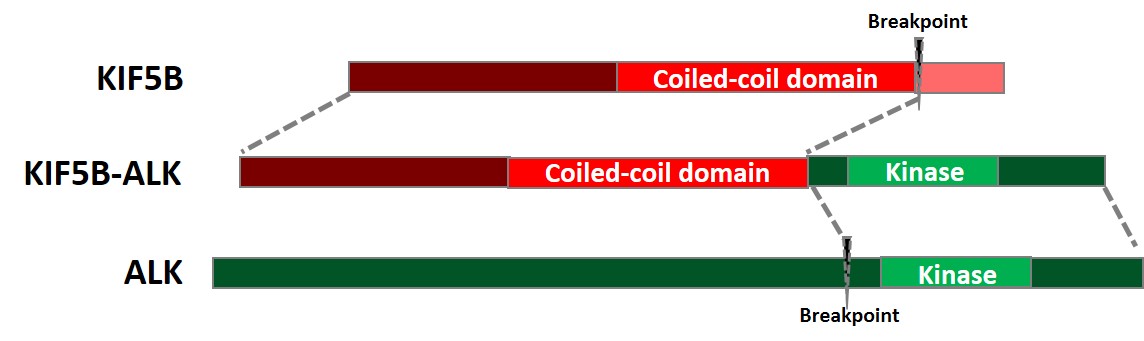

Figure 1. Schematic representation of the KIF5B[24]-ALK[19] gene fusion at the transcriptional level.
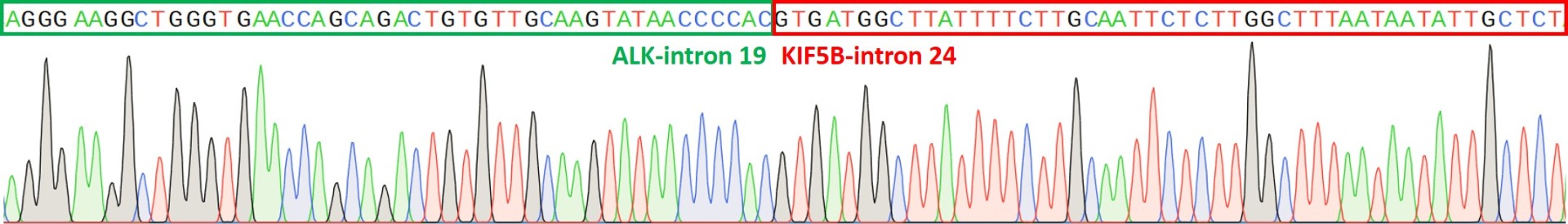
Figure 2. Representative sequence chromatogram of a clone containing the KIF5B-ALK fusion mutation in hiPSC, ASE-9211. The fusion mutation was engineered using proprietary designing strategies involving the co-transfection of guide RNA (gRNAs) and Cas9 to bind and cut the targeted intronic regions of the KIF5B and ALK genes. Single cell clones were shown confirmed to have the mutation by Sanger sequencing.
Application Notes
Chromosomal rearrangements (translocation, inversion, deletion, or insertion) of independent (separate) genes result in chimeric genes or gene fusions and have been implicated in many types of cancer (solid tumors and hematologic malignancies). Originally identified in the 1980s, these fusion genes have been identified to play a major role in precision/ personalized medicine, and are used as therapeutic biomarkers for diagnosis, prognosis, and drug development. However, the functional role of these mutations is not well understood. Most studies to elucidate mechanisms of the cancer mutations rely on gene knockout/ loss-of-function or insertion of recombinant gene and overexpression. However, these cell line models do not reflect the clinical situation of chromosomal translocation.
At Applied StemCell, we are experts in CRISPR/Cas9 cell line model generation and Induced Pluripotent Stem Cell (iPSC) technology. We provide custom service to engineer chromosomal rearrangement in your gene(s)-of-interest and iPSC line-of-choice that are most representative of your clinical samples:
- Two types of chimeric genes: at chromosomal level and chimeric cDNA knock-in
- Differentiation in T cell and other lineages
- Isogenic cell lines for reliable comparison of result and to elucidate full potential of mutations
- Multiple analysis at DNA, RNA and protein levels
- Detect with RT-PCR, RNA-Seq, Immunocytochemistry, Fluorescence in situ hybridization, and Western Blot
Applications
- Modeling for clinically relevant mutations
- Molecular biomarkers for cancer prognosis research
- Target for drug discovery and screening
- Immunotherapy research
Support Materials
Brochure/flyer:
Webinar:




