Newsletter
TARGATT™ HEK293 Master Cell Line For Site-Specific Knock-in
TARGATT™ HEK293 Master Cell Line and Knockin Kit - A valuable research tool to generate stable knock-in cell lines!
The TARGATT™ Knockin Master Cell Line and Kit uses integrase-based integration of a transgene into a preselected intergenic and transcriptionally active genomic locus (hROSA26, hH11, hAAVS1 or other safe harbor loci) engineered with an integrase recognition “attP” docking site or “landing-pad"). Applied StemCell (ASC) provides landing-pad ready TARGATT™ master cell lines and kits.
The TARGATT™ HEK293 Master Cell Line and Knockin Kit includes a TARGATT™ cloning plasmid that contains an integrase-recognition “attB” sequence and can be used to generate the donor plasmid containing the gene of interest (transgene). When the donor plasmid is transfected into the master cell line along with the integrase expression plasmid (also provided in the kit), the integrase catalyzes the integration of the transgene at the attP-attB sites. This integration is unidirectional which results in a stable integrated knock-in cell line.
The landing pad in the TARGATT™ HEK293 master cell line is engineered into the well-defined, transcriptionally active, intergenic H11 locus (safe harbor locus/genomic hotspot). This locus enables the high-level expression of the integrated gene-of-interest without disruption of internal genes and gene silencing commonly seen with random integration.
Advantages of the TARGATT™ HEK293 Master Cell Line:
- High knock-in efficiency
- Site-specific integration into the H11 genomic hotspot well-defined safe harbor locus
- Single gene knockin: one variant - one locus - one cell line
- Unidirectional integration for stable knock-in cell lines
- Uniform, high-level gene expression
- Overcomes challenges such as random insertion, gene silencing, multiple copy gene integration, ablated gene expression.
The TARGATT™ HEK293 Master Cell Lines and Knockin Kit combines the scalability, affordability, and ease of use of bacterial/yeast systems and the advantages of using mammalian cells (closer to the human environment and post-translational modifications) for efficient and stable gene knockin into cell lines and for library generation. If you would like to learn more, explore our product pages below (AST-1305) or contact us today!
If you would like to use your own cell line, our team can engineer the TARGATT™ system into your cell line of choice.
Products and Services
Technical Details

Schematic Representation of the Transgene Integration in the TARGATT™ Master Cell Line
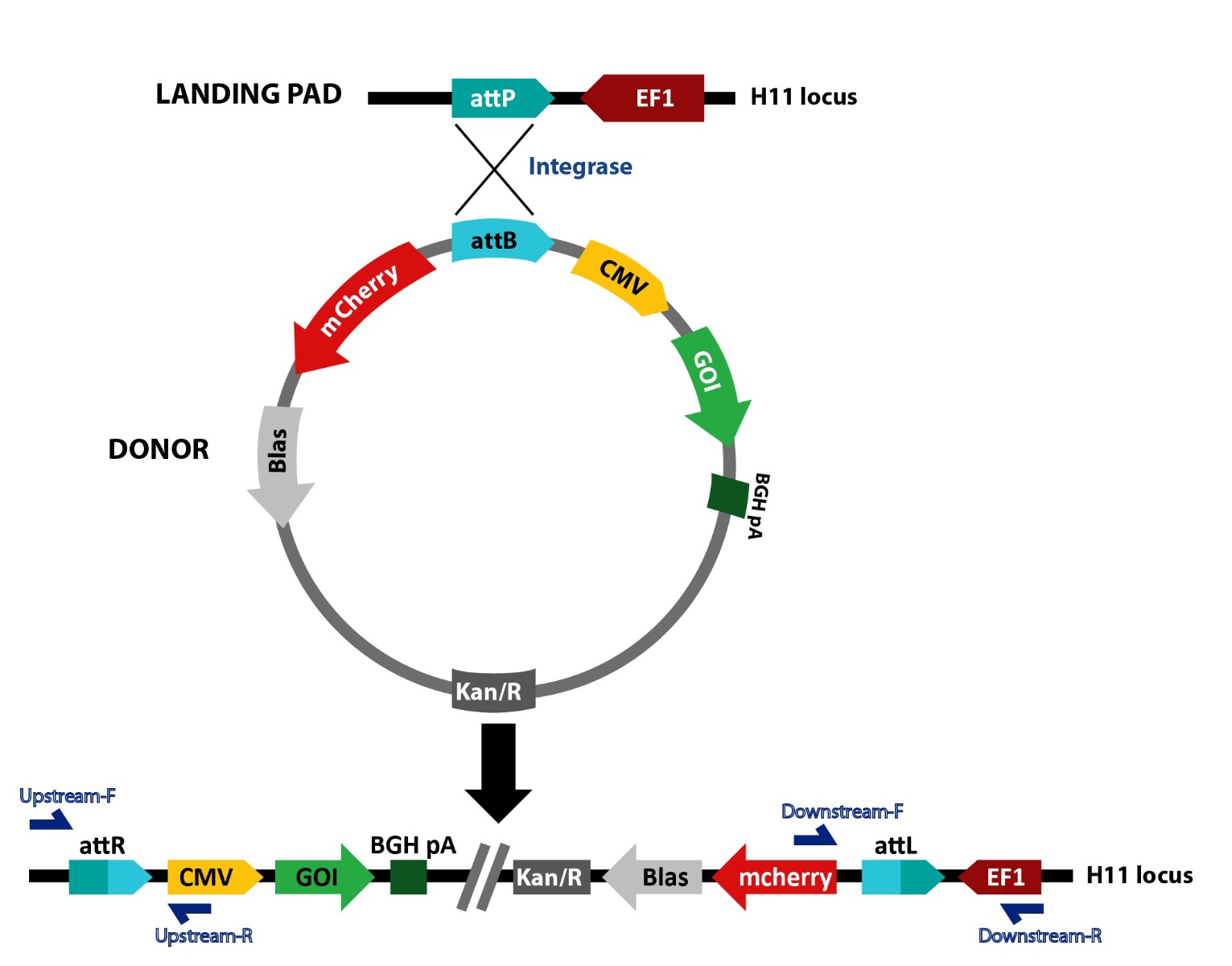
Figure 1. Schematic representation of TARGATT™ site-specific transgene integration mediated by integrase. The TARGATT™-HEK Master Cell Line was engineered with the attP landing pad at the hH11 safe harbor locus. The TARGATT™ plasmid containing the integrase recognition site, attB is used to clone the transgene. The integrase catalyzes an irreversible reaction between the attP site in the genome and attB site in the donor vector, resulting in integration of the gene of interest at the selected H11 locus. The cells containing the gene of interest can be enriched using the selection marker (gray box).
Confirmation of Site-Specific CMV-MCS Plasmid Integration
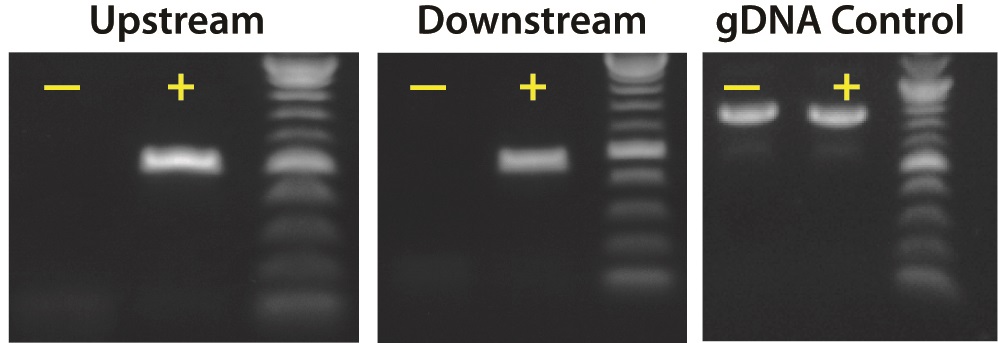
Figure 2. PCR gel electrophoresis to confirm the knockin of TARGATT™ 24 CMV-MCS-attB plasmid mediated by the TARGATT™ Integrase plasmid, after transfection into the TARGATT™ HEK293 Master Cell Line. Two sets of primers were used to confirm knockin: Upstream (512 bp) and Downstream primers (464 bp). The Human control primers (777 bp) was also used as a control to check the integrity of the cells and the genomic DNA (gDNA). Negative control (-) represents cells transfected with the TARGATT™ 24 CMV-MCS-attB plasmid and a mutant TARGATT™ integrase plasmid that is deficient for integration.
mCherry Expression After Transfection and Blasticidin Enrichment
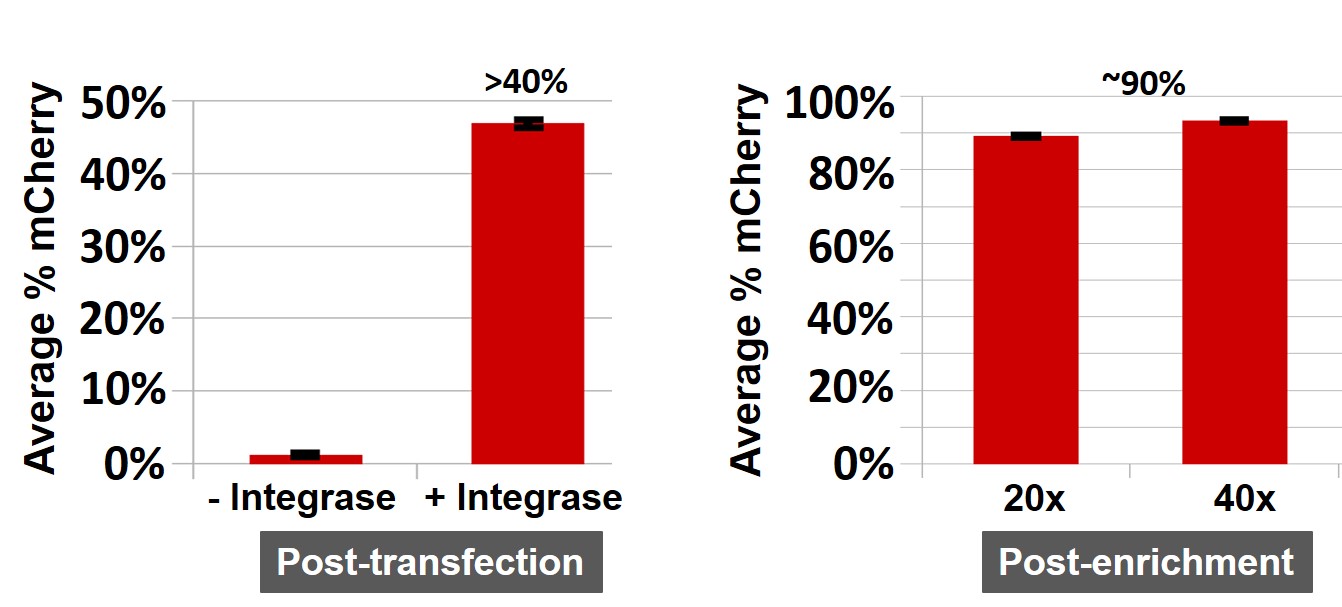
Figure 3. The mCherry integration into the TARGATT™ HEK293 master cell line. Left: Integration mediated by the integrase 72 hours post-transfection. Cells were transfected with the mCherry positive control plasmid and either the provided TARGATT™ integrase plasmid (+Integrase) or a mutant TARGATT™ integrase plasmid deficient for integration (-Integrase). The mCherry plasmid has no promoter and requires the ubiquitous EF1 promoter in the landing pad after integration to express the reporter gene. The integration efficiency of mCherry knockin into landing pad was >40%, without selection. Right: Blasticidin enrichment of TARGATT™ HEK293 cells with a knocked-in mCherry-blasticidin plasmid. Cell pools (with 20x and 40x split ratio) were enriched in selection medium for 3 weeks (without cell sorting). The enrichment of mCherry was about 90% after blasticidin selection. Data represents the mean ± SE of two representative experiments done in triplicates.
Comparing TARGATT™ and Existing Gene Editing Technologies For Generation Stable Knock-in Cell Lines
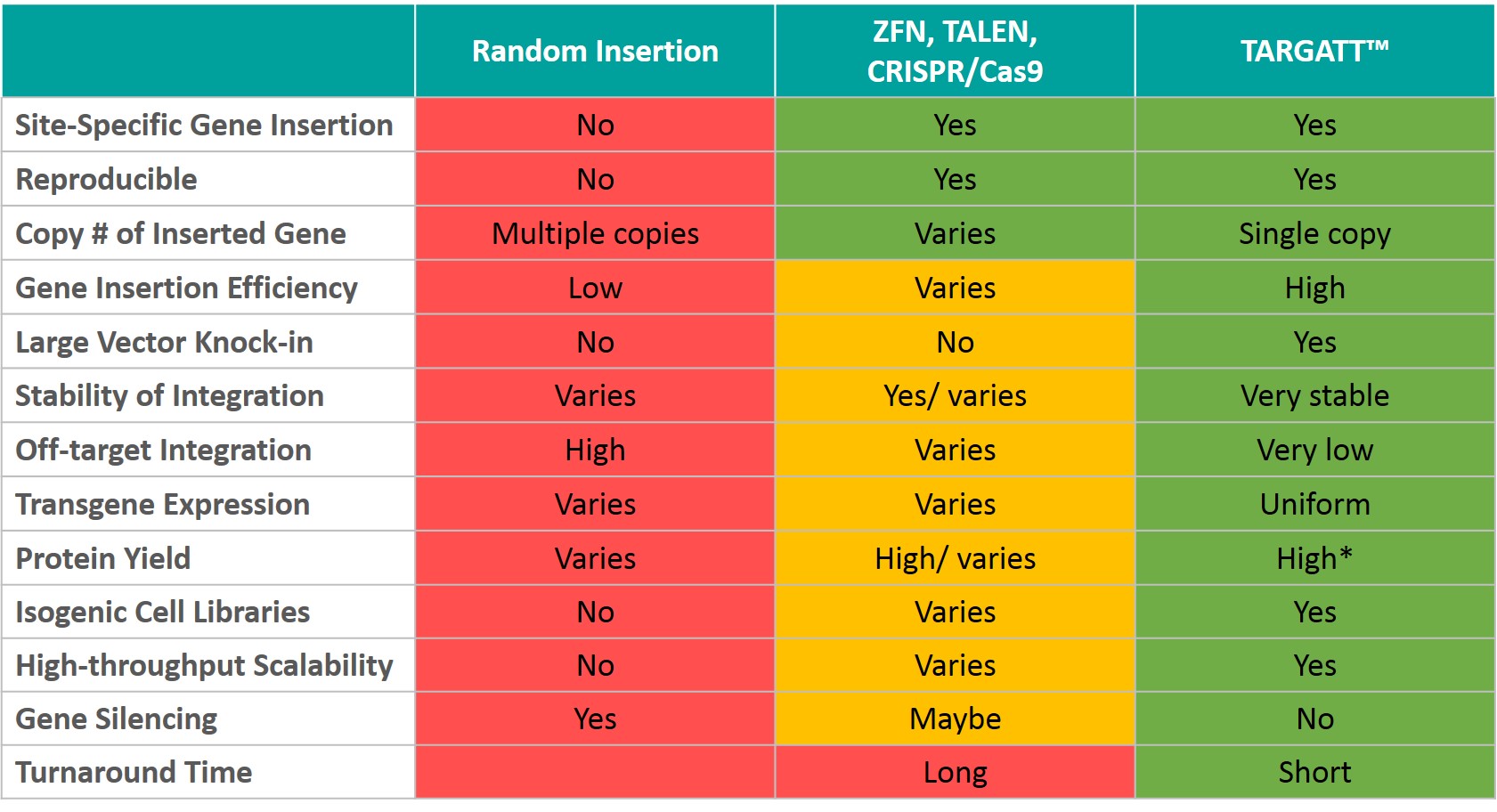
Please Note: The TARGATT™ integrase-based knockin technology requires the master cell line to have a landing pad (docking site) engineered into the cell at the chosen locus to be able to knockin efficiently. We have master cell lines in HEK293, CHO, and hiPSC backgrounds. We can also custom engineer a landing pad and generate a Master Cell Line in the cell line of your choice. Please inquire for further details.
How is the TARGATT™ HEK293 Master Cell Line Different From Other Technologies For Purpose of Site-Specific Integration of Transgenes and Stable Cell Line Generation?
The TARGATT™ integrase system is designed based on serine integrases which catalyze irreversible site-specific transgene insertion via recombination between two distinct integrase-recognition/ attachment sites, attP and attB. The difference of the TARGATT™ HEK293 Master Cell Line is its high efficiency in gene insertion (up to 45% compared to 5% using other systems) and its specificity with very low off-target integration profiles.
Another advantage of the TARGATT™ HEK293 Master Cell Line is its fast Blasticidin enrichment and mCherry selection (2-3 days vs weeks) that allows you to purify the gene edited cells most effectively.
How do the TARGATT™ HEK293 Master Cell Lines compare to other HEK293 Master Cell Lines and Kits Using Similar Site-Specific Technologies?
- High basal integration efficiency (up to 40%) even without selection or enrichment. With selection, modified cells can be enriched up to or higher than 90%.
- Unidirectional recombination ensures stable knock-in cell lines
- Single-step transfection procedure – no need for re-targeting
- Stringent and low off-target profile for random integration
- Integration locus, H11 is a well defined, intergenic, and transcriptionally active safe-harbor locus
- The H11 locus expresses high level of protein, uniformly and consistently
- Enables the generation of isogenic cell lines efficiently
- With selection/enrichment, stable-line pools can be obtained within 2 weeks.
How does TARGATT™ technology Compare To Nuclease-Based (Cas9) and other systems such as Flp, Cre, etc.?
The TARGATT™ technology is based on serine integrases which catalyze efficient site-specific DNA cleavage, DNA strand exchange and ligation without help from outside proteins. This permits knockin efficiencies superior to those that are possible with nucleases like Cas9, which leave cleaved DNA to be dealt with by the host repair machinery.
Bi-directional recombinases such as Flp and Cre also permit avoidance of the cell DNA repair machinery but are highly inefficient for net-integration due to the extreme kinetic favorability of the excision reaction. I.e., while they can mediate the initial genomic-recombination event with the same efficiency, they then proceed to catalyze the excision reaction with high efficiency (as the substrates are now physically linked). Serine integrase attL and attR complexes do not synapse, so this subsequent excision is blocked, and thus a high net-integration efficiency can be achieved.
You can also review the table in the Application Notes section for more comparisons between the TARGATT™ and other commonly used technologies.
Application Notes
Potential Applications include but are not limited to:
|
Immuno-oncology
|
Antibody Discovery
|
|
Protein evolution
|
|
|
Stem Cell Research
|
|
Support Materials
eBook:
TARGATT™ Technology For Antibody Discovery and Screening
(March 2021)
*Featured in Informa Connect's eBook: Antibody
Discovery, Selection & Screening
Poster:
![]() TARGATT™: Rapid and Efficient Transgene Integration Technology to Develop Large Mammalian Cell-Based Screening Libraries
TARGATT™: Rapid and Efficient Transgene Integration Technology to Develop Large Mammalian Cell-Based Screening Libraries
Webinar:
![]() TARGATT™ Rapid and Efficient Transgene Integration Technology: Stable Cell Line Development and Generation of Large Mammalian Cell-based Screening Libraries
TARGATT™ Rapid and Efficient Transgene Integration Technology: Stable Cell Line Development and Generation of Large Mammalian Cell-based Screening Libraries
Publications
TARGATT™ Master Cell Line
- Chi, X., Zheng, Q., Jiang, R., Chen-Tsai, R. Y., & Kong, L. J. (2019). A system for site-specific integration of transgenes in mammalian cells. PLOS ONE, 14(7), e0219842.
Transgenic Mouse Book Chapters
- Chen-Tsai, R. Y. (2020). Integrase-Mediated Targeted Transgenics Through Pronuclear Microinjection. In Transgenic Mouse (pp. 35-46). Humana, New York, NY.
- Chen-Tsai, R. Y. (2019). Using TARGATT™ Technology to Generate Site-Specific Transgenic Mice. In Microinjection (pp. 71-86). Humana Press, New York, NY.
Description of the technology
- Zhu, F., Gamboa, M., Farruggio, A. P., Hippenmeyer, S., Tasic, B., Schüle, B., … Calos, M. P. (2014). DICE, an efficient system for iterative genomic editing in human pluripotent stem cells. Nucleic Acids Research, 42(5), e34. http://doi.org/10.1093/nar/gkt1290.
- Tasic, B., Hippenmeyer, S., Wang, C., Gamboa, M., Zong, H., Chen-Tsai, Y., & Luo, L. (2011). Site-specific integrase-mediated transgenesis in mice via pronuclear injection. Proceedings of the National Academy of Sciences of the United States of America, 108(19), 7902–7907. http://doi.org/10.1073/pnas.1019507108.
Commentary, comparison with other transgenic methods
- Rossant, J., Nutter, L. M., & Gertsenstein, M. (2011). Engineering the embryo. Proceedings of the National Academy of Sciences, 108(19), 7659-7660.
Tet inducible mice generated by TARGATT™
- Fan, X., Petitt, M., Gamboa, M., Huang, M., Dhal, S., Druzin, M. L., … Nayak, N. R. (2012). Transient, Inducible, Placenta-Specific Gene Expression in Mice. Endocrinology, 153(11), 5637–5644. http://doi.org/10.1210/en.2012-1556.
Advantage of Hipp11 (H11) locus
- Hippenmeyer, S., Youn, Y. H., Moon, H. M., Miyamichi, K., Zong, H., Wynshaw-Boris, A., & Luo, L. (2010). Genetic Mosaic Dissection of Lis1 and Ndel1 in Neuronal Migration. Neuron, 68(4), 695–709. http://doi.org/10.1016/j.neuron.2010.09.027.
Applications for TARGATT™ technology
- Lindtner, S., Catta-Preta, R., Tian, H., Su-Feher, L., Price, J. D., Dickel, D. E., ... & Pennacchio, L. A. (2019). Genomic Resolution of DLX-Orchestrated Transcriptional Circuits Driving Development of Forebrain GABAergic Neurons. Cell reports, 28(8), 2048-2063.
- Wang, T. A., Teo, C. F., Åkerblom, M., Chen, C., Tynan-La Fontaine, M., Greiner, V. J., ... & Jan, L. Y. (2019). Thermoregulation via Temperature-Dependent PGD2 Production in Mouse Preoptic Area. Neuron, 103(2), 309-322.
- Clarke, B. A., Majumder, S., Zhu, H., Lee, Y. T., Kono, M., Li, C., ... & Byrnes, C. (2019). The Ormdl genes regulate the sphingolipid synthesis pathway to ensure proper myelination and neurologic function in mice. eLife, 8.
- Carlson, H. L., & Stadler, H. S. (2019). Development and functional characterization of a lncRNA‐HIT conditional loss of function allele. genesis, e23351.
- Chande, S., Ho, B., Fetene, J., & Bergwitz, C. (2019). Transgenic mouse model for conditional expression of influenza hemagglutinin-tagged human SLC20A1/PIT1. PloS one, 14(10), e0223052. doi:10.1371/journal.pone.0223052
- Hu, Q., Ye, Y., Chan, L. C., Li, Y., Liang, K., Lin, A., ... & Pan, Y. (2019). Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nature immunology, 1.
- Matharu, N., Rattanasopha, S., Tamura, S., Maliskova, L., Wang, Y., Bernard, A., ... & Ahituv, N. (2018). CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science, eaau0629.
- Chen-Tsai, R. Y. (2019). Using TARGATT™ Technology to Generate Site-Specific Transgenic Mice. In Microinjection (pp. 71-86). Humana Press, New York, NY
- Barrett, R. D., Laurent, S., Mallarino, R., Pfeifer, S. P., Xu, C. C., Foll, M., ... & Hoekstra, H. E. (2018). The fitness consequences of genetic variation in wild populations of mice. bioRxiv, 383240.
- Ibrahim, L. A., Huang, J. J., Wang, S. Z., Kim, Y. J., Li, I., & Huizhong, W. (2018). Sparse Labeling and Neural Tracing in Brain Circuits by STARS Strategy: Revealing Morphological Development of Type II Spiral Ganglion Neurons. Cerebral Cortex, 1-14.
- Kumar, A., Dhar, S., Campanelli, G., Butt, N. A., Schallheim, J. M., Gomez, C. R., & Levenson, A. S. (2018). MTA 1 drives malignant progression and bone metastasis in prostate cancer. Molecular oncology.
- Jang, Y., Broun, A., Wang, C., Park, Y. K., Zhuang, L., Lee, J. E., ... & Ge, K. (2018). H3. 3K4M destabilizes enhancer H3K4 methyltransferases MLL3/MLL4 and impairs adipose tissue development. Nucleic acids research. https://doi.org/10.1093/nar/gky982
- Tang, Y., Kwon, H., Neel, B. A., Kasher-Meron, M., Pessin, J., Yamada, E., & Pessin, J. E. (2018). The fructose-2, 6-bisphosphatase TIGAR suppresses NF-κB signaling by directly inhibiting the linear ubiquitin assembly complex LUBAC. Journal of Biological Chemistry, jbc-RA118.
- Chen, M., Geoffroy, C. G., Meves, J. M., Narang, A., Li, Y., Nguyen, M. T., ... & Elzière, L. (2018). Leucine Zipper-Bearing Kinase Is a Critical Regulator of Astrocyte Reactivity in the Adult Mammalian CNS. Cell Reports, 22(13), 3587-3597
- Kido, T., Sun, Z., & Lau, Y.-F. C. (2017). Aberrant activation of the human sex-determining gene in early embryonic development results in postnatal growth retardation and lethality in mice. Scientific Reports, 7, 4113. http://doi.org/10.1038/s41598-017-04117-6.
- Nouri, N., & Awatramani, R. (2017). A novel floor plate boundary defined by adjacent En1 and Dbx1 microdomains distinguishes midbrain dopamine and hypothalamic neurons. Development, 144(5), 916-927.
- Li, K., Wang, F., Cao, W. B., Lv, X. X., Hua, F., Cui, B., ... & Yu, J. M. (2017). TRIB3 Promotes APL Progression through Stabilization of the Oncoprotein PML-RARα and Inhibition of p53-Mediated Senescence. Cancer Cell, 31(5), 697-710.
- Jiang, T., Kindt, K., & Wu, D. K. (2017). Transcription factor Emx2 controls stereociliary bundle orientation of sensory hair cells. eLife, 6, e23661.
- Booze, M. L., Hansen, J. M., & Vitiello, P. F. (2016). A Novel Mouse Model for the Identification of Thioredoxin-1 Protein Interactions. Free Radical Biology & Medicine, 99, 533–543. http://doi.org/10.1016/j.freeradbiomed.2016.09.013.
- Feng, D., Dai, S., Liu, F., Ohtake, Y., Zhou, Z., Wang, H., ... & Hayat, U. (2016). Cre-inducible human CD59 mediates rapid cell ablation after intermedilysin administration. The Journal of clinical investigation, 126(6), 2321-2333.
- Sun, N., Yun, J., Liu, J., Malide, D., Liu, C., Rovira, I. I., … Finkel, T. (2015). Measuring in vivo mitophagy. Molecular Cell, 60(4), 685–696. http://doi.org/10.1016/j.molcel.2015.10.009.
- Devine, W. P., Wythe, J. D., George, M., Koshiba-Takeuchi, K., & Bruneau, B. G. (2014). Early patterning and specification of cardiac progenitors in gastrulating mesoderm. eLife, 3, e03848. http://doi.org/10.7554/eLife.03848.
- Fogg, P. C. M., Colloms, S., Rosser, S., Stark, M., & Smith, M. C. M. (2014). New Applications for Phage Integrases. Journal of Molecular Biology, 426(15), 2703–2716. http://doi.org/10.1016/j.jmb.2014.05.014.
- Chen-Tsai, R. Y., Jiang, R., Zhuang, L., Wu, J., Li, L., & Wu, J. (2014). Genome editing and animal models. Chinese science bulletin, 59(1), 1-6.
- Park, K.-E., Park, C.-H., Powell, A., Martin, J., Donovan, D. M., & Telugu, B. P. (2016). Targeted Gene Knockin in Porcine Somatic Cells Using CRISPR/Cas Ribonucleoproteins. International Journal of Molecular Sciences, 17(6), 810. http://doi.org/10.3390/ijms17060810.
- Guenther, C. A., Tasic, B., Luo, L., Bedell, M. A., & Kingsley, D. M. (2014). A molecular basis for classic blond hair color in Europeans. Nature Genetics, 46(7), 748–752. http://doi.org/10.1038/ng.2991.
- Villamizar, C. A. (2014). Characterization of the vascular pathology in the acta2 r258c mouse model and cerebrovascular characterization of the acta2 null mouse. UT GSBS Dissertations and These (Open Access). Paper 508 (2014)
FAQs
How did you confirm a single landing pad, was it copy number qPCR?
What is the safe-harbor H11 Locus?
Are kit 1305 and 1306 using the same cells TARGATT HEK293 (AST1305-1)?
Can you share the HEK293 suspension culture protocol? So that we could evaluate whether it is possible for us to make library cells in suspension condition before we commercialize it.
Does TARGATT™ 24 CMV-MCS cloning plasmid (AST-3064) express mCherry after recombination?
Could you explain a bit more about why AST-1306 won't express target after transfection? It seems SV40 promoter is already there in donor plasmid.
Once I purchase/license in the kit, are we able to modify the library plasmid by ourselves?
Do you share the sequence of the AST-3064 cloning plasmid used in the AST-1305 kit?
What is the antibiotic resistance marker to amplify the AST-3065 mCherry positive control and AST-3201 integrase Plasmid?
Can I use DH5-alpha competent cells for transforming the donor plasmid instead of NEB® 10-beta cells?
Can the AST-3064 TARGATT™ cloning plasmid be used for generating mammalian cell libraries with the TARGATT™ HEK293 Master Cell Line?
How can I design primers to confirm insertion of the gene of interest for the TARGATT™ HEK293 Master Cell Line?
Do you have TARGATT™ Master Cell Lines in other cell line backgrounds in addition to the TARGATT™ HEK293 Master Cell Line?
Can I use other TARGATT™ Plasmids such as AST-3050 or AST-3051 with the TARGATT™ HEK293 Master Cell Line?
Can I use Hygromycin or G418 in my cell culture system?
We also would like to use hygromycin to select for TARGATT cells inserted with a hygromycin resistance gene. On the website, it says we cannot use hygromycin on TARGATT HEK. Why is that?
Will the integration of the plasmid backbone into the TARGATT™ HEK293 Master Cell Line affect expression of the gene of interest (transgene)?
Can you generate a TARGATT™ Master Cell Line with the cell line I am interested?
Can you generate a TARGATT™ Master Cell Line with docking site at a different locus?
What is the size of the transgene that can be integrated into the TARGATT™ HEK293 Master Cell Line with the specified efficiency?
AST-1305: I ordered your kit AST-1305. In 5.4 of the manual, after 72 hours transfection, is it OK to plant the 20 fold diluted cells in blasticidin selection medium directly without growth medium first then medium replacement after 24 hours.
AST-1305: Does the landing pad have both attP and EF1 promoter? Or only attP? If EF1 a promoter is present, how big is the promoter?
AST-1305: Does the landing pad only integrate into one of the 2 chromosomes? How did you verify it is a single copy instead of 2 copies?
AST-1305: How is the plasmid for library cloning different from AST-3064? Does it still have mCherry and Blasticidin resistant gene? Is the GOI still under CMV promoter? Also, does it have an ampicillin marker?
AST-1305: Do you provide a marker to detect transfection efficiency?
AST-1305: Can this kit be used to build libraries?
AST-1305: When are mCherry and blas expressed?
AST-1305: Could you please share the protocols for cloning of monocistronic (single gene) and bicistronic (IgG) constructs?
Are the TARGATT™ HEK293 cells adherent?
Do you have data that shows the inserted copy number is actually a single copy?
Could you let me know the ratio of the single copy number/ total inserted copy number?
Which HEK293 cell line is the original Cell Line for this kit (there are some kind of type (e.g. HEK293T, HEK293S)?
Could let me know the efficiency of insertion with genes that have 20 kb and 10 kb lengths respectively?
After the transgene integration, the attR and attL sequence are left completely or sometimes have indels?
Can you confirm whether attP and attB sites are always divided at the same sequence part?
If I would like to use a different promotoer, what should I do?
Does the AST-3064 plasmid that have a blast gene too in addition to the mCherry? If AST-3064 already has mCherry, why do you also provide AST-3065?
What is the difference between AST-3064, AST-3065, and AST-3075?