Newsletter
Reporter iPSC Lines (isogenic)
Applied StemCell has engineered two isogenic panels of reporter gene iPSC lines from a fully characterized parental control/ master line (ASE-9109). These reporter lines are ideal for quantitative investigation of intricate genetic and molecular mechanisms underlying lineage-specific development, disease pathology and drug interactions.
-
Two Isogenic panels of “knock-in” (KI) reporter iPSCs engineered from fully characterized parental control iPSCs:
-
Neuronal Lineage Specific Reporter iPSCs
-
Safe Harbor Locus Reporter (Master) iPSCs
-
Fully characterized reporter iPSCs: pluripotency, differentiation capability, and functionality
-
Demonstrated use of these reporter lines for high throughput neurotoxicity screening of drug candidates
For custom reporter line generation, please contact us.
Products and Services
Technical Details
Genome engineering in induced pluripotent stem cells (iPSCs) offers the advantage of generating numerous cell line models in an isogenic background along with the parental cell line as an ideal control, where patient-derived primary cells are difficult to obtain. As well, genome editing to introduce/ correct mutations and knock-in reporter genes in control and disease iPSC lines, is a powerful tool for comparative study of genetic mechanisms involved in cellular/organ development and disease pathology. Furthermore, the directed differentiation of iPSCs into a desired cell line lineage such as neuronal lineage cells, provides a consistent and predictive source of disease models.
Applications:
-
Research models to quantitatively study genetic and molecular mechanisms during development, differentiation and disease pathogenesis
-
Quantitative assessment of genetically modified cells in cell-based therapies
Preclinical drug development and screening applications (drug screening of neuroprotective and neurotoxicological compounds)
|
1. Lineage-specific Reporter Gene Knock-in |
|
Lineage-specific Reporters |
Description |
| MAP2-Nanoluc-Halotag KI | Neuron reporter |
| GFAP-Nanoluc-Halotag KI | Astrocyte reporter |
| MBP-Nanoluc-Halotag KI | Oligodendrocyte reporter |
|
2. Safe Harbor Locus Reporter Gene Knock-in iPSCs: |
|
Safe-harbor Reporters |
Description |
| CAG-GFP, AAVS/Chr19 | Ubiquitous reporter |
| DCXp-GFP | Neuron reporter |
Application Notes
Lineage-specific Reporter Knock-in iPSC Line: Neuronal Lineage-specific Line (ASE-9500)
The ASE-9500 iPSC line is a mono-allelic (heterozygous) knock-in iPSC line engineered with a multiplexed reporter (Nanoluc-Halotag) inserted into the 3’ end of the endogenous MAP2 locus. The parental iPSC is ASE-9109 which is an integration-free, normal karyotype iPSC derived from male, CD34+ cord blood cells. The MAP2-Nanoluc-Halotag Knock-in iPSC line can be further differentiated into an isogenic panel of neurons and glia for disease modeling and drug/ toxicity screening applications.
1. Schematic representation of MAP2-Nanoluc-Halotag reporter construct
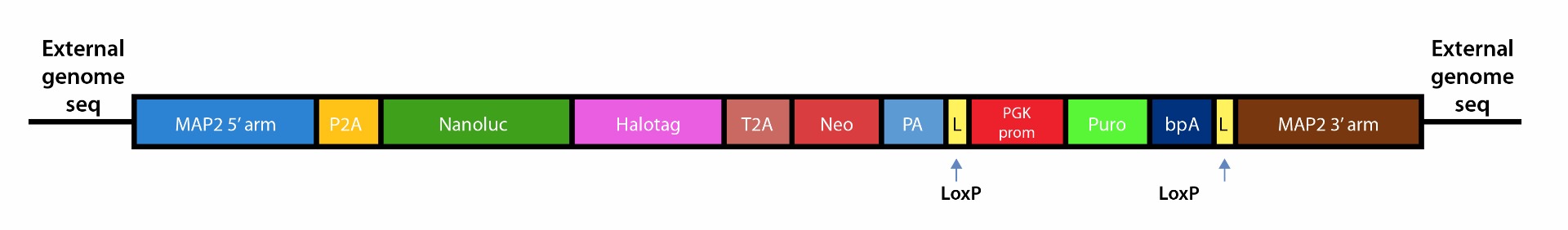
Figure 1. The dual reporter, Nanoluc-Halotag construct is expressed by neuronal lineage-specific promoter, MAP2. This construct is inserted into the endogenous MAP2 locus of the iPSC line.
2. Validation of MAP2-Nanoluc-Halotag knock-in clone by PCR
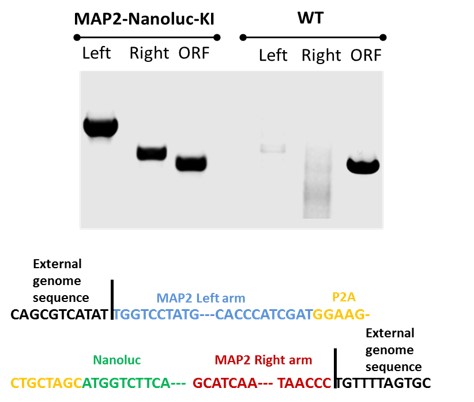
Figure 2. PCR was used to confirm the insertion of reporter genes to the MAP2 locus in ASE-9500 MAP2-Nanoluc-Halotag knock-in (KI) heterozygous iPSC line. Three sets of primers were used to verify insertion of the reporter gene at the MAP2 locus: 3’, right arm integration; 5’, left arm integration; wild type (WT) open reading frame (ORF) confirmed reporter gene KI at the correct locus. Sequence validation for ASE-9500 MAP2-Nanoluc-Halotag knock-in (KI) heterozygous iPSC line.
3. Pluripotency marker validation and karyotype analysis of MAP2-Nanoluc-Halotag reporter knock-in iPSC line
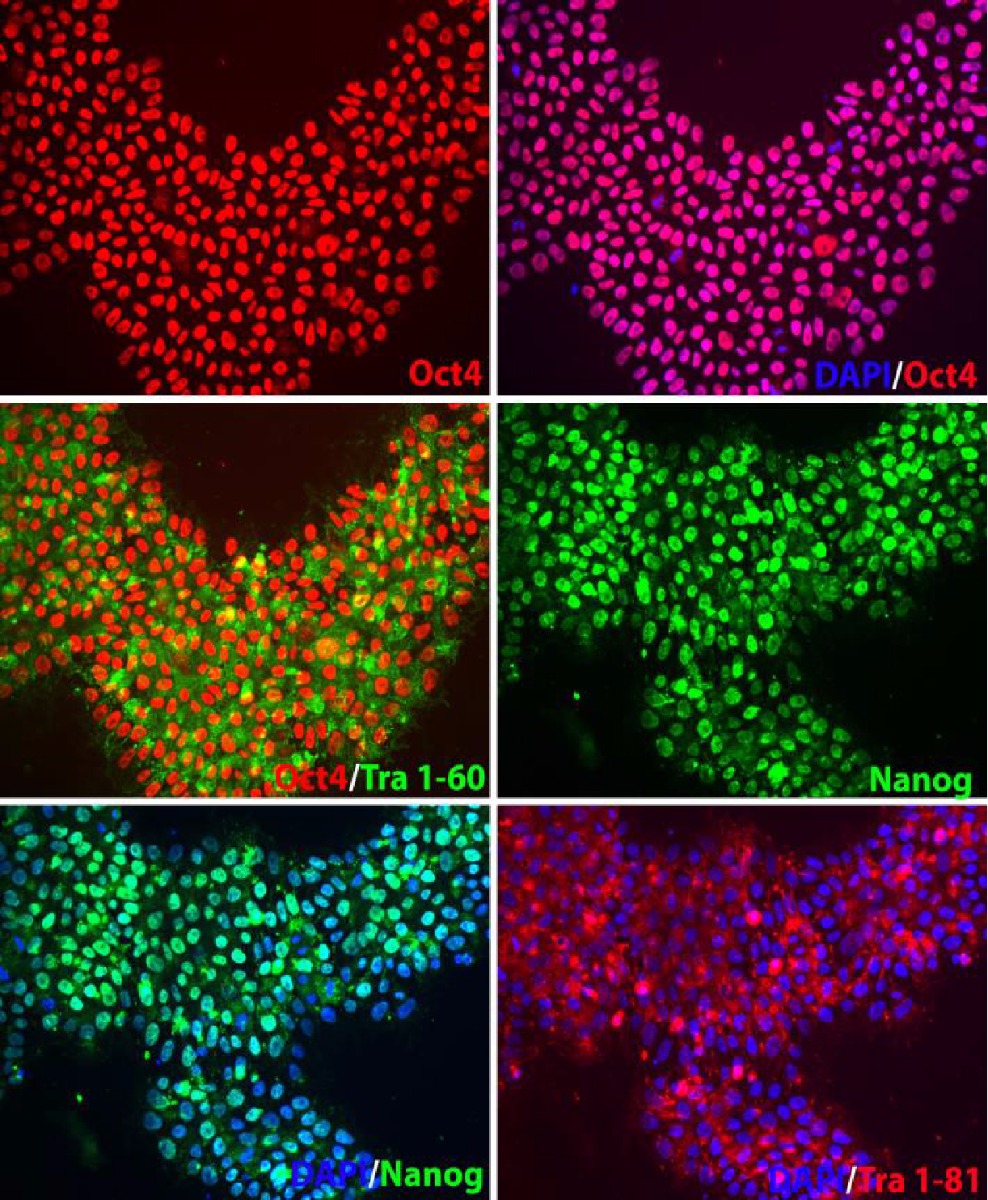
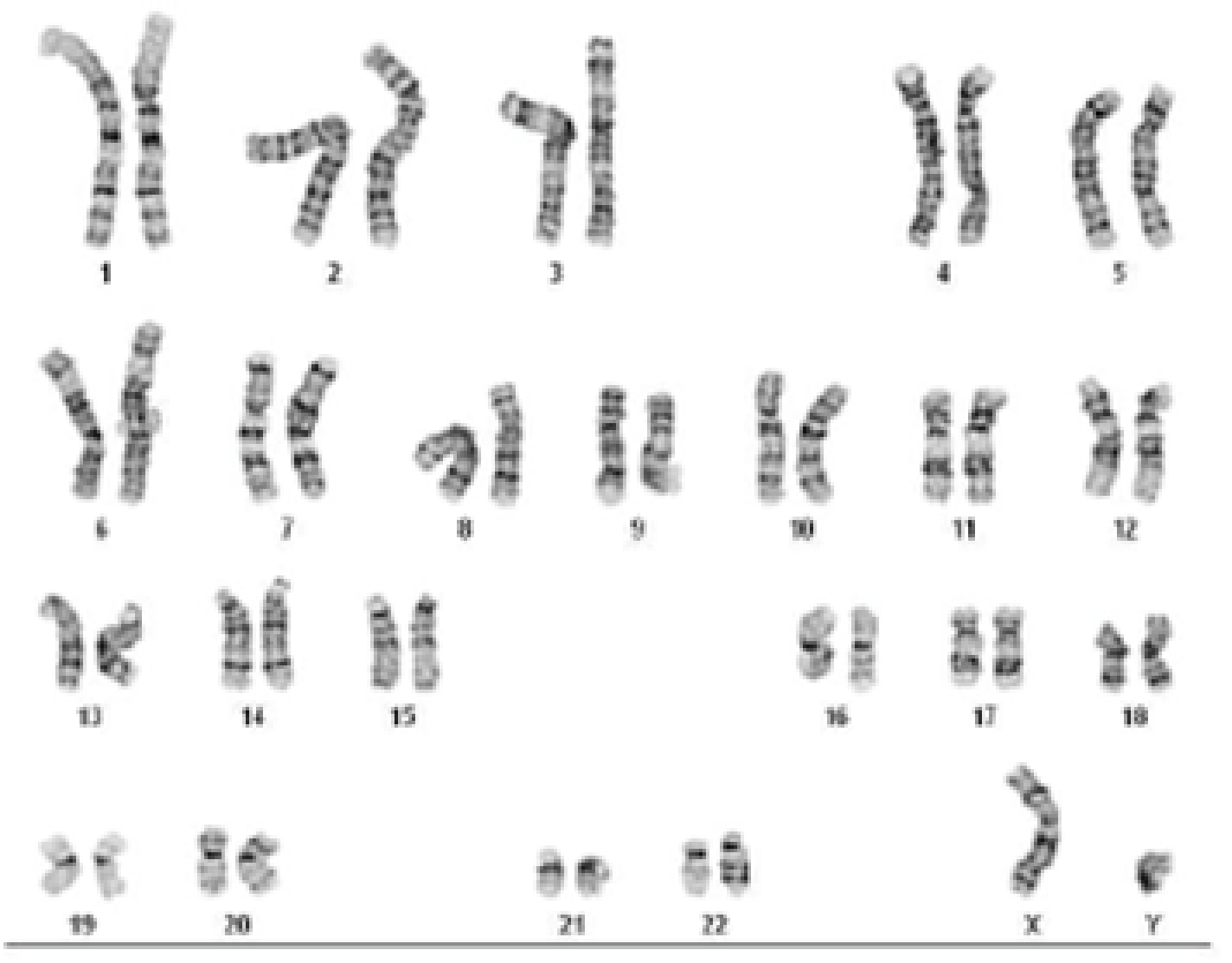
Figure 3. Pluripotency characterization and karyotype analysis of ASE-9500 Lineage-specific reporter knock-in iPSC line, MAP2-Nanoluc-Halotag heterozygous knock-in line. The MAP2-Nanoluc-Halotag line expresses pluripotency markers OCT4, NANOG, TRA-1-60, and TRA-1-81, indicating pluripotency of the iPSC line after genome editing. Nucleus stained with DAPI (blue). Karyotype Analysis of MAP2-Nanoluc-Halotag iPSC line indicates normal karyotype and no clonal abnormalities in the iPSC line at the stated band level of resolution, after genome editing.
4. Functional validation of neuronal lineage-specific reporter line, ASE-9500 MAP2-Nanoluc-Halotag knock-in iPSC line.
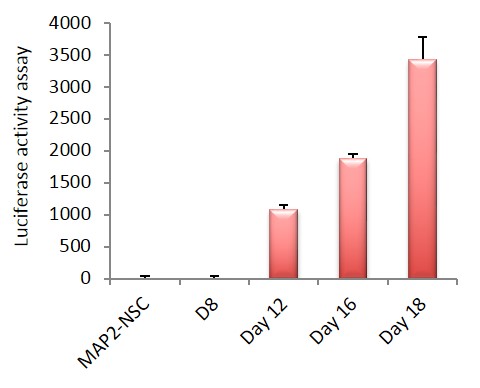
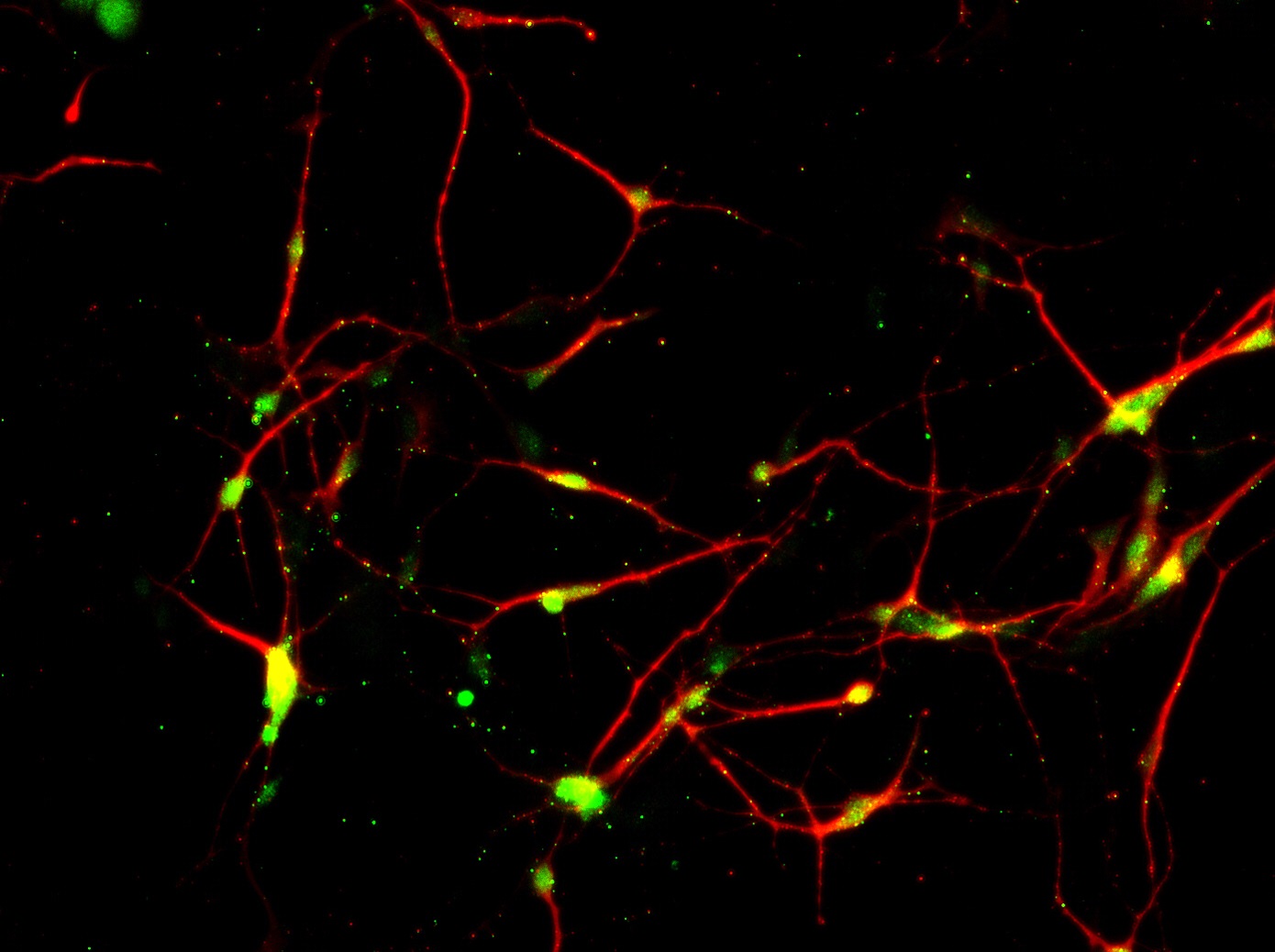
Figure 4. Functional validation of ASE-9500 Lineage-specific reporter knock-in iPSC line, MAP2-Nanoluc-Halotag heterozygous knock-in line by luciferase activity assay measured over 18 days from the neural stem cell to neuronal maturation. Immunohistochemical staining for MAP2 (red) and Halotag (green) showed co-localization (yellow) of reporter gene in neuronal cells.
iPSC lines with reporter gene inserted at a safe harbor locus: AAVS1-DCXp-GFP iPSC line (ASE-9502)
ASE-9502 iPSC line is a mono-allelic (heterozygous) knock-in iPSC line engineered with a GFP reporter tag driven by a neuronal-lineage-specific promoter, Doublecortin (DCX) inserted into the AAVS safe harbor locus. The parental iPSC is ASE-9109 which is an integration-free, normal karyotype iPSC derived from male, CD34+ cord blood cells. The AAVS-DCX-GFP Knock-in iPSC line can be further differentiated into an isogenic panel of neurons and glia for disease modeling and drug/ toxicity screening applications.
1. Schematic representation of AAVS-DCXp-GFP reporter construct
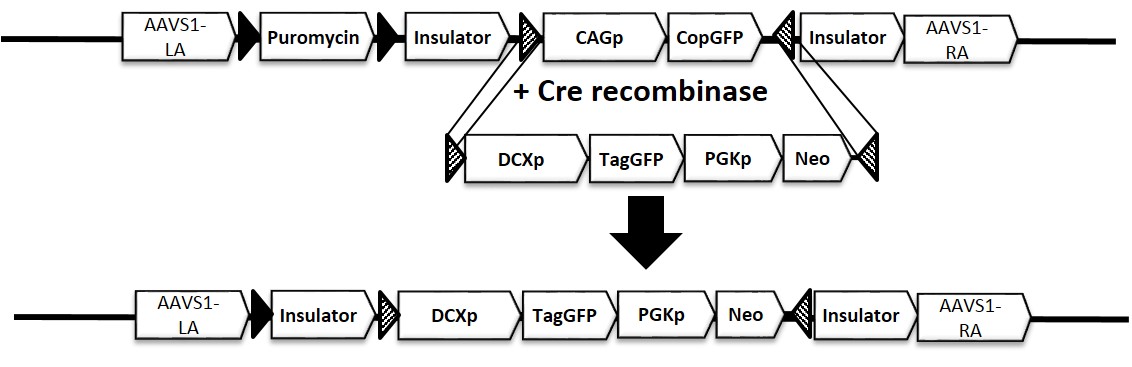
Figure 1. Schematic representation of iPSC line with reporter gene inserted in AAVS safe harbor locus, AAVS1-DCXp-GFP (ASE-9502). The reporter gene expression is driven by a neuronal-lineage (neuronal development) specific promoter, Doublecortin (DCXp). The ASE-9502 cell line is derived from master cell line with a ubiquitous promoter expressing a CopGFP reporter gene. After Cre-recombinase mediated cassette exchange (RMCE), a single copy of the DCXp-TagGFP cassette is inserted into the AAVS1 locus.
2. Validation of neuronal-lineage specific reporter cassette knock-in in AAVS1 safe harbor locus
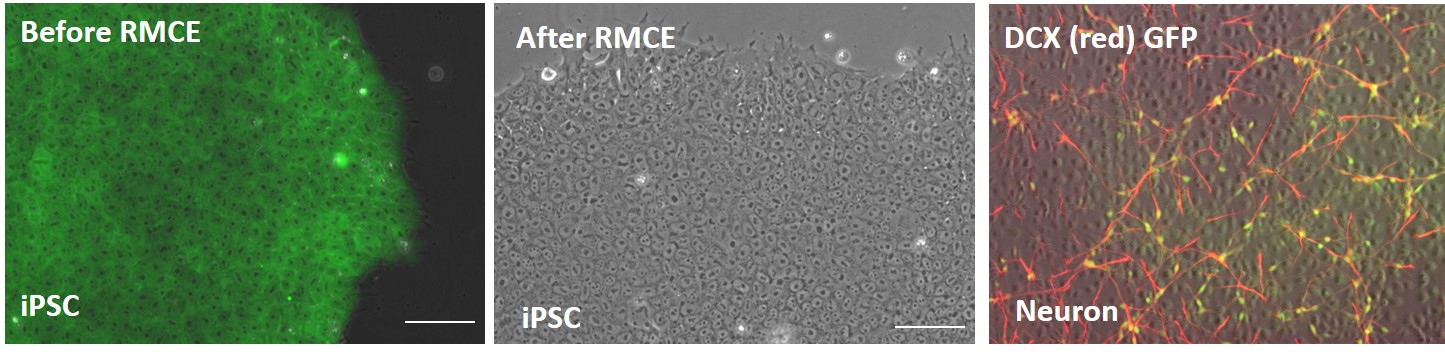
Figure 2. Validation of the DCXp-GFP reporter gene insertion in AAVS1 locus in iPSC line, ASE-9502. In the AAVS-CAG-GFP master cell line, GFP expression is driven by the ubiquitous promoter, CAG (all cells are stained green for GFP). After RMCE, there is no GFP expression in the iPSCs confirming the absence of a ubiquitous CAG promoter after cassette exchange. When these iPSCs are differentiated into neurons, immunohistochemical characterization showed the co-localization (yellow) of GFP (green) and DCXp-positive neurons (red). The GFP is expressed only in neuronal cells confirming the insertion of the reporter gene in the correct locus.
Support Materials
Brochure/Flyer:
 iPSC Generation, Disease Modeling, Mutation Correction and Differentiation
iPSC Generation, Disease Modeling, Mutation Correction and Differentiation iPSC-based CNS Drug Efficacy and Neurotoxicity Testing
iPSC-based CNS Drug Efficacy and Neurotoxicity Testing Genome Editing for Cell Line Model Generation
Genome Editing for Cell Line Model Generation ASC's Custom Services for Cell Line and Animal Model Generation, and Downstream Assays for Drug/ Toxicity Testing
ASC's Custom Services for Cell Line and Animal Model Generation, and Downstream Assays for Drug/ Toxicity TestingPosters:
FAQs
How specific is the GFP reporter driven by the DCXp promoter in the AAVS1-DCXp-GFP knock-in iPSC line?
Do you observe a loss/decrease of reporter activity in the astrocyte lineage-specific reporter iPSC line (ASE-9501; GFAP-Nanoluc-Halotag) and other reporter iPSC lines, as the cells differentiate?