WGS Data is Now Available Upon Request - Inquire
MTA required
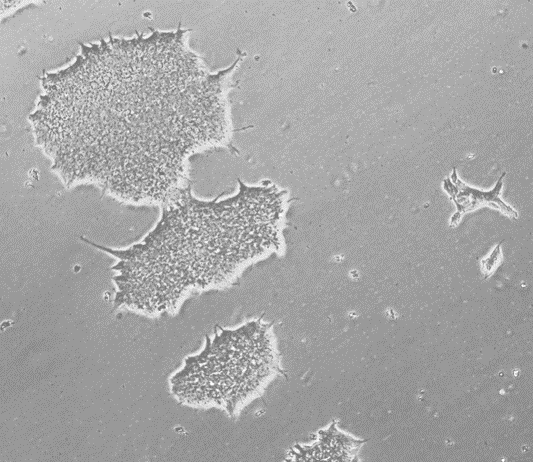
Applied StemCell (ASC), Inc. provides Control Human Induced Pluripotent Stem Cells (iPSC) at low passages (p15). These pluripotent cells were generated from normal human skin fibroblasts using episomal reprogramming methods. This method allows the transient expression of human transcription factors (OCT4, SOX2, KLF4, and c-MYC) that initiate the reprogramming process. The resulting human iPS cells (hiPSCs) were selected using morphological criteria without the use of either fluorescent markers or drug selection. These iPS cells have been tested for the expression of the pluripotency markers, including OCT4, SOX2, SSEA4, TRA-1-60, TRA-1-81, and alkaline phosphatase (AP) activity, and normal male karyotype. The ASE-9211 control human iPSC line has proven capability for CRISPR/Cas9 genome editing and differentiation to various somatic lineages.
*Detailed protocols for thawing, culturing under feeder-free conditions, and cryopreservation of these iPS cells are provided. For donor information, please refer to the datasheet or contact us.
Applied StemCell has developed an efficient integration-free, small molecule-based method to differentiate a variety of high-quality cells from the ASE-9211 iPSC Control Master Cell Line.
iPSC-Derived Cells:
- Natural Killer Cells
- T Cells
- Human Retinal Pigment Epithelium (RPE)
- Human Hematopoietic Progenitor Cell (HPC)
- Photoreceptor Cells
- Myoblast
- Cardiomyocytes
If you do not see what you are looking for or you would like to differentiate your healthy or diseases iPSCs, please visit our iPSC Differentiation Service page or contact us today.
Characterization
Pluripotency Marker Analysis
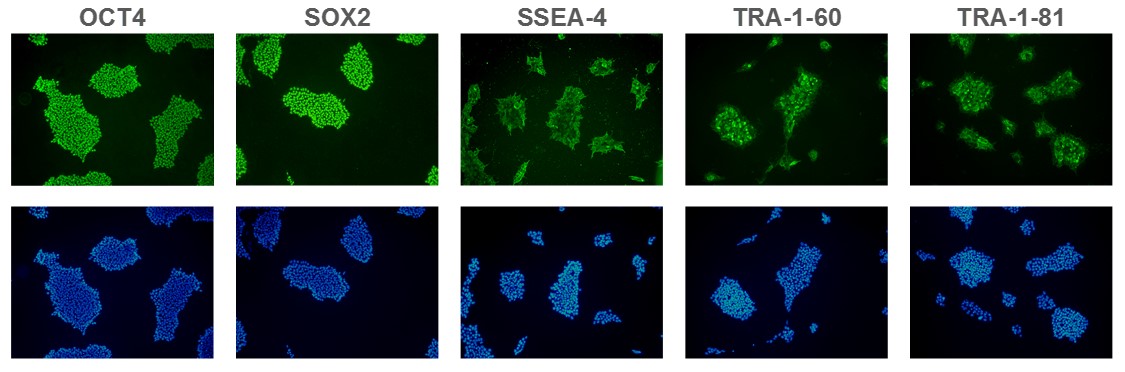
Figure 1. Expression of pluripotency markers. ASC 9211 iPS cell line expresses common iPSC biomarkers (top row: OCT-4, SOX2, SSEA-4, TRA-1-60, and TRA-1-81). Bottom row: Corresponding DAPI nuclear staining. All images were taken at 10x magnification.
Alkaline Phosphatase (AP) Staining
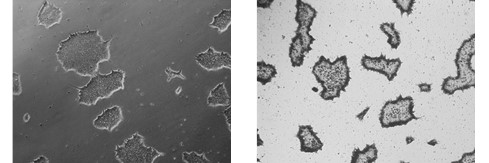
Figure 2. Alkaline Phosphatase (AP) staining. ASC-9211 iPSCs stain positive for Alkaline Phosphatase: a typical unstained colony (a) was used to gauge the extent of the AP staining (b). Both images were taken at 5x magnification.
Karyotype Analysis
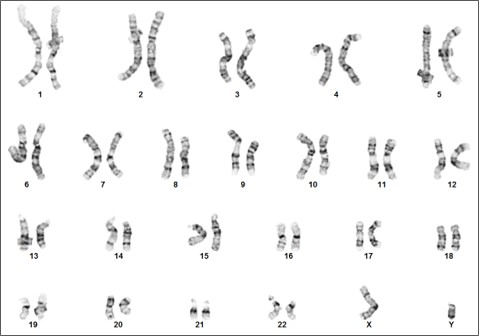
Figure 3. Karyotype analysis to rule out genetic aberrations. This iPSC line demonstrates a normal male karyotype. Cytogenic analysis was performed on twenty G-banded metaphase cells from human iPSC line, ASE-9211 at passage 15. Nineteen cells demonstrated an apparently normal male karyotype, and one cell demonstrated a non-clonal chromosome aberration, which is most likely an artifact of culture.
Karyotype Analysis
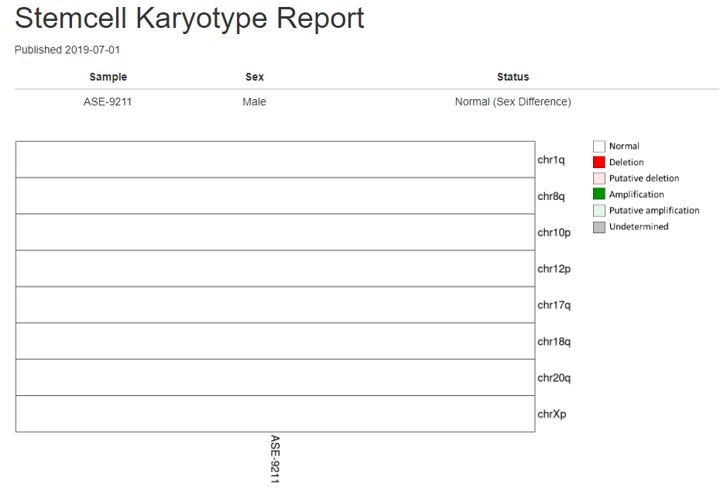
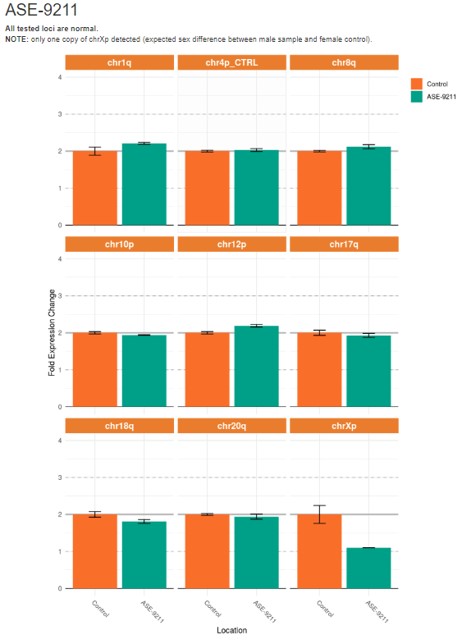
Figure 4. Quantitative PCR (qPCR) analysis to detect karyotypic abnormalities in control human iPSC line, ASE-9211 shows a normal male karyotype in all loci tested.
Directed-differentiation to Three Germ Layers
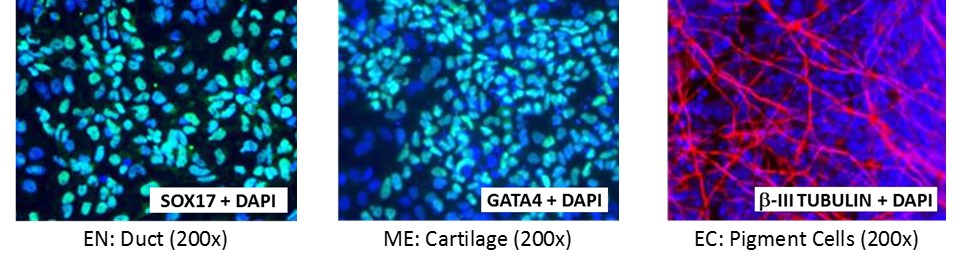
Figure 5. Direct differentiation of ASE-9211 p18 to three germ layers. Immunofluorescent staining for lineage-specific biomarkers of three germ layers after direct differentiation of control hiPSC line, ASE-9211. The ASE-9211 hiPSC line was differentiated to specific lineages of the germ layers using well-established and optimized protocols. Immunostaining for biomarkers of each lineage was performed to confirm lineage commitment. Cells were also co-stained with nuclear marker, DAPI (blue). Images shows are co-localization of biomarker with DAPI. Endoderm (EN) marker: SOX17 (green); Mesoderm (ME) marker: GATA4 (green); Ectoderm (EC) marker: b-III Tubulin (red).
Teratoma Formation Pluripotency Analysis Report
Identification of germ layers from the kidney and testis, and tumors derived from NST-R2 cell lines (H&E staining)
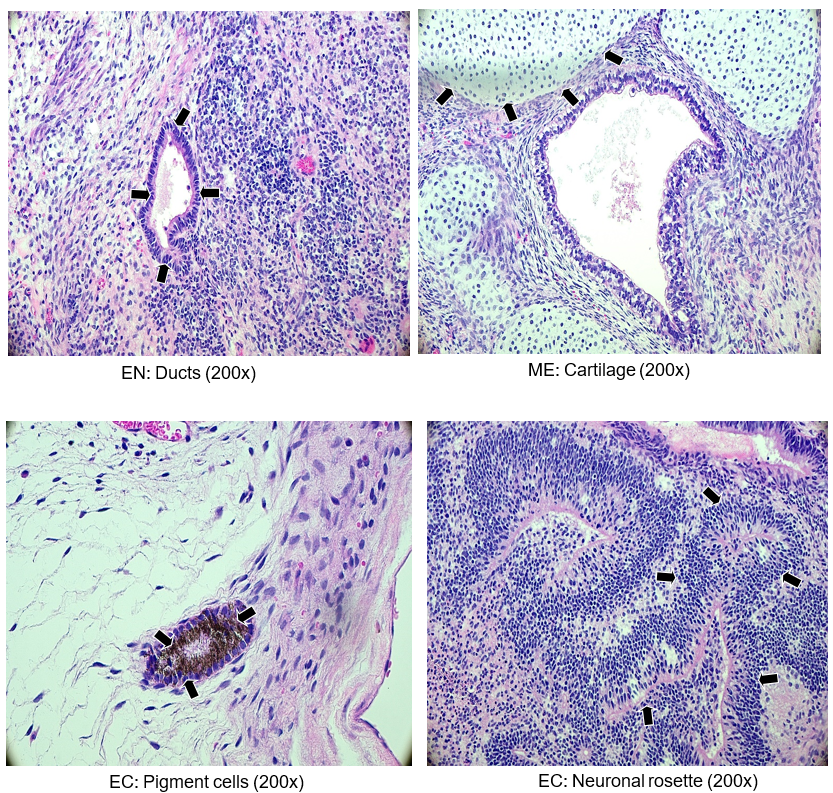
Figure 6. Histological analyses of kidney and testis injected with cell line ASE-9211. Variously differentiated tissues that represent the three germ layers are shown and indicated by arrow heads. Abbreviations: EN, endoderm, ME, mesoderm, EC, ectoderm
CRISPR Genome Engineering Capability

Figure 6. CRISPR/Cas9 was used to generate 6 knockout cell lines from the control human iPSC line, ASE-9211.
Differentiation Capability
Control iPSC line, ASE-9211 was used for differentiation into motor neurons using proprietary, integration-free protocols.
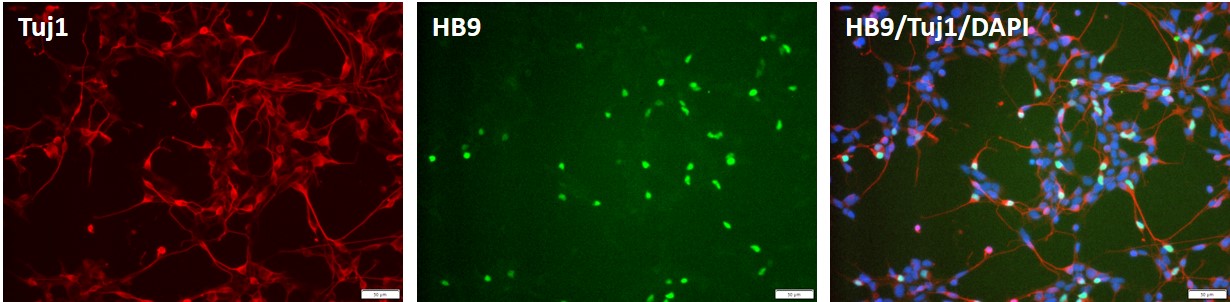
Figure 7. Immunocytochemical staining images of the motor neuron derived from control "master" iPSC line, ASE-9211 at 2 days post thaw. The iPSC-derived motor neurons were stained with antibodies for motor neuron biomarker, HB9 (green), and neuronal biomarker, Tuj1 (red). DAPI (blue) was used as nucleus stain.
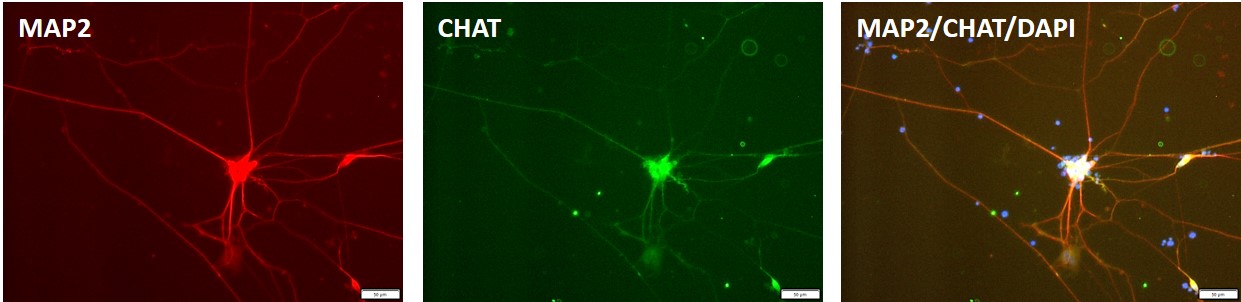
Figure 8. Immunocytochemical staining for motor neuron marker, ChAT (green) and neuronal marker, MAP2 i(blue) n motor neurons differentiated from control "master" iPSC line, ASE-9211 at 7 days post thaw. DAPI (blue) was used for nucleus staining.